
Alternative Estimation Frameworks with `mars`
mars
authors
2026-05-15
Alternative-Estimation-Frameworks.RmdThis vignette demonstrates three alternative estimation frameworks
available in mars:
- Bootstrap
- Jackknife (leave-one-cluster-out for multilevel models)
- Permutation
For each framework, we show how to obtain:
- Point estimate from the empirical distribution of replicate estimates
- 95% interval (
2.5%,97.5%) from that same distribution
library(mars)
extract_alt_interval <- function(x, term = "(Intercept)") {
if (!is.null(x$summary) &&
all(c("point_estimate", "ci_lower", "ci_upper") %in% colnames(x$summary)) &&
term %in% rownames(x$summary)) {
out <- x$summary[term, c("point_estimate", "ci_lower", "ci_upper"), drop = FALSE]
return(out)
}
if (!is.null(x$summary) &&
all(c("estimate", "ci_lower", "ci_upper") %in% colnames(x$summary)) &&
term %in% rownames(x$summary)) {
out <- x$summary[term, c("estimate", "ci_lower", "ci_upper"), drop = FALSE]
names(out) <- c("point_estimate", "ci_lower", "ci_upper")
return(out)
}
if (!is.null(x$quantiles) &&
all(c("2.5%", "50.0%", "97.5%") %in% colnames(x$quantiles)) &&
term %in% rownames(x$quantiles)) {
out <- x$quantiles[term, c("50.0%", "2.5%", "97.5%"), drop = FALSE]
names(out) <- c("point_estimate", "ci_lower", "ci_upper")
return(out)
}
if (!is.null(x$summary) &&
all(c("estimate", "jackknife_se") %in% colnames(x$summary)) &&
term %in% rownames(x$summary)) {
est <- x$summary[term, "estimate"]
se <- x$summary[term, "jackknife_se"]
out <- data.frame(
point_estimate = est,
ci_lower = est - 1.96 * se,
ci_upper = est + 1.96 * se,
row.names = term
)
return(out)
}
stop("Could not extract a point estimate + interval for term: ", term)
}Data and Base Model
We use the bundled school dataset and a multilevel
model.
fit <- mars(
formula = effect ~ 1 + (1 | district/study),
data = school,
studyID = "district",
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE"
)
summary(fit)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multilevel
#>
#> Estimation Method:
#> Maximum Likelihood
#>
#> Model Formula:
#> effect ~ 1 + (1 | district/study)
#>
#> Data Summary:
#> Number of Effect Sizes: 56
#> Number of Fixed Effects: 1
#> Number of Random Effects: 2
#>
#> Random Data Summary:
#> Number unique district: 11
#> Number unique study: 56
#>
#> Random Components:
#> term var SD
#> district 0.05774 0.2403
#> study 0.03286 0.1813
#>
#> Fixed Effects Estimates:
#> attribute estimate SE z_test p_value lower upper
#> (Intercept) 0.1845 0.08048 2.292 0.02191 0.02671 0.3422
#>
#> Model Fit Statistics:
#> logLik Dev AIC BIC AICc
#> -8.395 24.79 22.79 28.87 24.5
#>
#> Q Error: 578.864 (55), p < 0.0001
#>
#> I2 (General):
#> names values
#> district 92.38
#> study 87.34
#>
#> I2 (Jackson): 98.69621
#> I2 (Between): 86.3727
#>
#> Residual Diagnostics:
#> n n_finite_raw mean_raw sd_raw rmse mae q_pearson mean_abs_studentized
#> 56 56 -0.06446 0.3258 0.3293 0.2632 96.8 1.065
#> max_abs_studentized prop_abs_studentized_gt2 prop_abs_studentized_gt3
#> 4.196 0.1071 0.03571
#>
#> Normality (whitened residuals): test n_tested statistic p_value
#> shapiro_wilk_whitened 56 0.9555 0.03766
#>
#> Heteroscedasticity trend (|raw residual| ~ fitted): n corr_abs_raw_fitted slope p_value
#> 56 NA NA NABootstrap
set.seed(2026)
boot_repl <- replicate_bootstrap(
number_bootstraps = 5,
data = school,
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE"
)
boot_stats <- compute_alt_stats(boot_repl, type = "bootstrap")The summary component contains distribution-based point
estimates and 95% intervals:
summary(boot_stats)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multilevel
#>
#> Estimation Method:
#> MLE
#>
#> Robust Method:
#> Bootstrap
#> Number of Replications: 5
#>
#> Model Formula:
#> effect ~ 1 + (1 | district/study)
#>
#> Data Summary:
#> Number of Effect Sizes: 56
#> Number of Fixed Effects: 1
#> Number of Random Effects: 2
#>
#> Random Components (Bootstrap CIs):
#> Variances:
#> 50% 2.5% 97.5%
#> Tau_1 0.0681 0.0163 0.0741
#> Tau_2 0.0223 0.0083 0.0252
#>
#> SDs:
#> 50% 2.5% 97.5%
#> district 0.2610 0.1257 0.2722
#> study 0.1492 0.0906 0.1587
#>
#> Fixed Effects Estimates (Bootstrap CIs):
#> 50% 2.5% 97.5%
#> (Intercept) 0.1519 0.1015 0.193
#>
#> Model Fit Statistics (Bootstrap CIs):
#> 50% 2.5% 97.5%
#> logLik -7.8490 -21.8355 6.7273
#> Dev 23.6981 -5.4547 51.6710
#> AIC 21.6981 -7.4547 49.6710
#> BIC 27.8272 -1.3474 55.7948
#> AICc 23.3781 -5.7601 51.3544The quantiles component provides named quantile columns
directly:
boot_stats$quantiles["(Intercept)", c("2.5%", "50.0%", "97.5%")]
#> 2.5% 50.0% 97.5%
#> 0.1014548 0.1519059 0.1930013All coefficient distributions can also be visualized in a single base R figure, with one histogram panel per effect:
plot(boot_stats)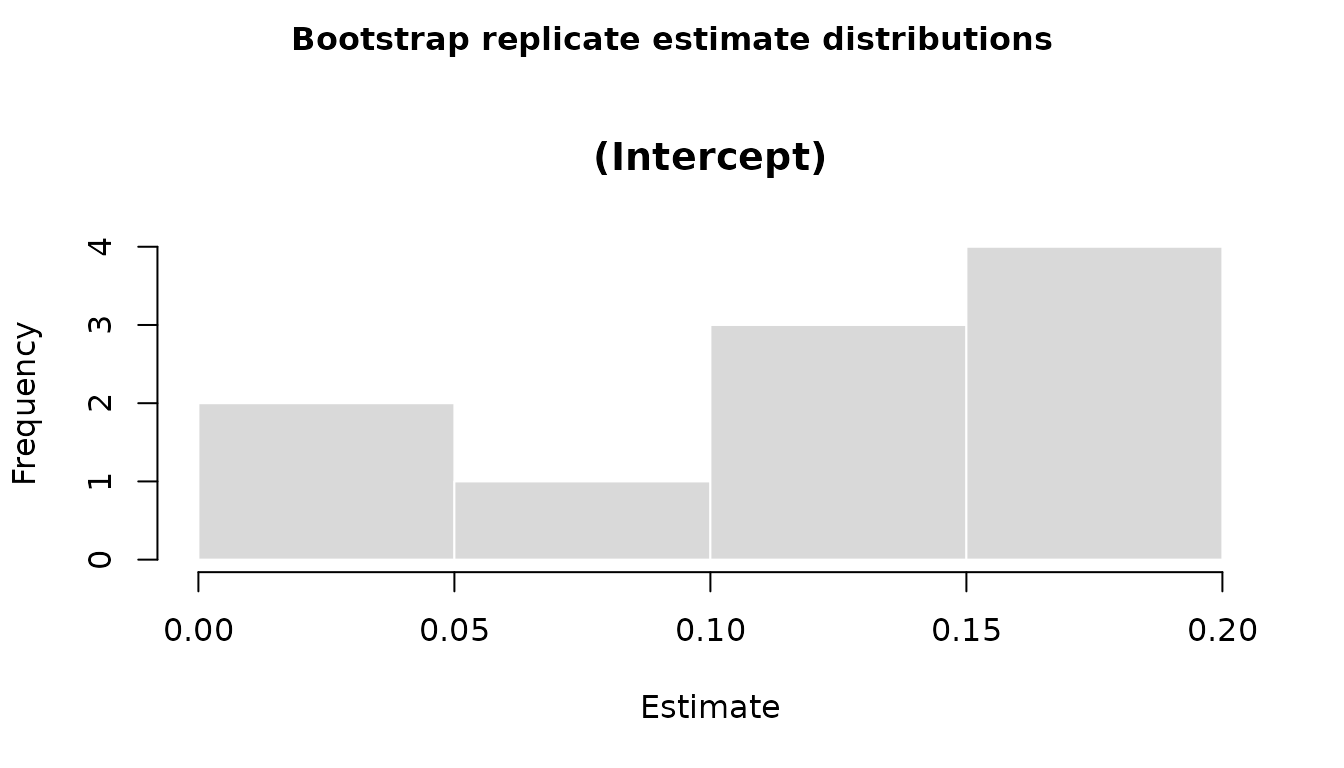
Jackknife
For multilevel models, jackknife leaves out one highest-level cluster
at a time (here: one district per replication). To keep the
vignette quick to compile, the example below uses only the first four
districts; the code is the same for the full dataset.
set.seed(2026)
school_jack <- subset(school, district %in% sort(unique(district))[1:4])
jack_repl <- replicate_jackknife(
data = school_jack,
structure = "multilevel",
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
estimation_method = "MLE"
)
jack_stats <- compute_alt_stats(jack_repl, type = "jackknife")Jackknife summaries include both the distribution-based interval and a classic jackknife standard error:
summary(jack_stats)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multilevel
#>
#> Estimation Method:
#> MLE
#>
#> Robust Method:
#> Jackknife
#> Number of Replications: 4
#>
#> Model Formula:
#> effect ~ 1 + (1 | district/study)
#>
#> Data Summary:
#> Number of Effect Sizes: 11
#> Number of Fixed Effects: 1
#> Number of Random Effects: 2
#>
#> Random Components (Jackknife CIs):
#> Variances:
#> 50% 2.5% 97.5%
#> Tau_1 0.0220 0.0123 0.0353
#> Tau_2 0.0273 0.0214 0.0368
#>
#> SDs:
#> 50% 2.5% 97.5%
#> district 0.1467 0.1107 0.1879
#> study 0.1646 0.1462 0.1919
#>
#> Fixed Effects Estimates (Jackknife CIs):
#> 50% 2.5% 97.5%
#> (Intercept) 0.2397 0.1457 0.3044
#>
#> Model Fit Statistics (Jackknife CIs):
#> 50% 2.5% 97.5%
#> logLik -0.9935 -2.3117 0.1567
#> Dev 9.9870 7.6866 12.6234
#> AIC 7.9870 5.6866 10.6234
#> BIC 9.1807 6.8802 12.0586
#> AICc 28.1921 26.6006 29.1758Permutation
For multilevel models, permutation shuffles highest-level clusters while keeping rows within each cluster together.
set.seed(2026)
perm_repl <- mars_alt_estimation(
type = "permutation",
number_permutations = 5,
data = school,
structure = "multilevel",
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
estimation_method = "MLE"
)
perm_stats <- compute_alt_stats(perm_repl, type = "permutation")Permutation uses the same summary structure as bootstrap:
summary(perm_stats)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multilevel
#>
#> Estimation Method:
#> MLE
#>
#> Robust Method:
#> Permutation
#> Number of Replications: 5
#>
#> Model Formula:
#> effect ~ 1 + (1 | district/study)
#>
#> Data Summary:
#> Number of Effect Sizes: 56
#> Number of Fixed Effects: 1
#> Number of Random Effects: 2
#>
#> Random Components (Permutation CIs):
#> Variances:
#> 50% 2.5% 97.5%
#> Tau_1 0.0577 0.0577 0.0577
#> Tau_2 0.0329 0.0329 0.0329
#>
#> SDs:
#> 50% 2.5% 97.5%
#> district 0.2403 0.2403 0.2403
#> study 0.1813 0.1813 0.1813
#>
#> Fixed Effects Estimates (Permutation CIs):
#> 50% 2.5% 97.5%
#> (Intercept) 0.1531 0.1147 0.1656
#>
#> Model Fit Statistics (Permutation CIs):
#> 50% 2.5% 97.5%
#> logLik -8.3949 -8.3949 -8.3949
#> Dev 24.7899 24.7899 24.7899
#> AIC 22.7899 22.7899 22.7899
#> BIC 28.8659 28.8659 28.8659
#> AICc 24.5042 24.5042 24.5042Compare the Three Frameworks
The example below extracts the intercept summaries side-by-side.
comparison <- rbind(
bootstrap = extract_alt_interval(boot_stats),
jackknife = extract_alt_interval(jack_stats),
permutation = extract_alt_interval(perm_stats)
)
comparison
#> point_estimate ci_lower ci_upper
#> bootstrap 0.1519059 0.1014548 0.1930013
#> jackknife 0.2397353 0.1457071 0.3043585
#> permutation 0.1531493 0.1146949 0.1655870In practice, increase the number of replications for stable inference (for example, 500 to 2000+), especially for interval estimation.