
Network Meta-analysis with MARS
mars authors
2026-05-15
Network-Meta-Analysis.RmdThis vignette shows how to run a network meta-analysis with
network_meta() using both multilevel and multivariate
fitting strategies, and how to extract:
- Direct effects
- Indirect effects
- Total effects
- Standard errors for direct, indirect, and total effects
- Normal-approximation confidence intervals for direct, indirect, and total effects
- Number of direct and indirect effects and contributing studies by comparison
- Heterogeneity summaries (
QandI^2, total and within) - Between-study variance (
tau^2) by model component/level - Incoherence-factor assessment (per-comparison and global)
- Contribution matrix (% information from direct comparisons)
- Treatment ranking/order and rank probabilities
- Node-splitting direct-vs-indirect comparisons
- Formatted league tables and prediction intervals
- Design-level inconsistency diagnostics
- Arm-level to contrast-level pairwise conversion
- Comparison-adjusted funnel plots and small-study asymmetry tests
- Fit-stat summaries and lightweight model comparison helper
library(mars)Example Network Data
Each row is an observed contrast with effect defined as
treatment_2 - treatment_1.
nma_dat <- data.frame(
study = c("s1", "s1", "s2", "s3", "s4", "s5", "s6", "s7", "s8", "s9", "s10", "s11"),
trt1 = c("A", "A", "A", "B", "A", "B", "C", "A", "B", "C", "A", "B"),
trt2 = c("B", "C", "B", "C", "D", "D", "D", "C", "D", "D", "B", "C"),
yi = c(0.22, 0.41, 0.18, 0.19, 0.61, 0.39, 0.23, 0.37, 0.43, 0.18, 0.24, 0.20),
vi = c(0.04, 0.05, 0.03, 0.04, 0.06, 0.05, 0.05, 0.04, 0.06, 0.05, 0.04, 0.04),
stringsAsFactors = FALSE
)
nma_dat
#> study trt1 trt2 yi vi
#> 1 s1 A B 0.22 0.04
#> 2 s1 A C 0.41 0.05
#> 3 s2 A B 0.18 0.03
#> 4 s3 B C 0.19 0.04
#> 5 s4 A D 0.61 0.06
#> 6 s5 B D 0.39 0.05
#> 7 s6 C D 0.23 0.05
#> 8 s7 A C 0.37 0.04
#> 9 s8 B D 0.43 0.06
#> 10 s9 C D 0.18 0.05
#> 11 s10 A B 0.24 0.04
#> 12 s11 B C 0.20 0.04To keep the vignette fast to compile, the live examples below use one
multilevel fit and one multivariate fit, both with moderators.
Additional model variants are shown later as eval = FALSE
templates.
Multilevel Network Meta-analysis
Multilevel network meta-analysis always estimates flexible
heterogeneity by random level. tau_components chooses the
component definition. This example also includes moderators so one
fitted object shows the full workflow.
nma_ml <- network_meta(
data = nma_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multilevel",
tau_components = "comparison",
moderators = ~ severity + followup_months,
ci_level = 0.90,
estimation_method = "MLE"
)
summary(nma_ml)
#> Network Meta-analysis (MARS)
#> Model Type: multilevel
#> Heterogeneity: flexible
#> Tau components: comparison
#> Moderators: severity, followup_months
#> Reference: A
#> Treatments: A, B, C, D
#>
#> Evidence Summary:
#> $n_comparisons_total
#> [1] 6
#>
#> $n_comparisons_with_direct
#> [1] 6
#>
#> $n_comparisons_with_indirect
#> [1] 6
#>
#> $n_comparisons_indirect_only
#> [1] 0
#>
#> $n_direct_effects
#> [1] 12
#>
#> $n_indirect_effects
#> [1] 48
#>
#>
#> Incoherence-Factor Assessment:
#> Global test:
#> Q_incoherence df p_value
#> 9.934809 6 0.1274243
#>
#> Per-comparison incoherence factors:
#> treatment_1 treatment_2 incoherence_factor if_se z p_value
#> A B 0.2682828 0.4318174 0.6212876 0.5344104
#> A C -0.3675848 0.3107142 -1.1830320 0.2367965
#> A D -0.5633491 0.3546417 -1.5885022 0.1121728
#> B C 0.1833121 0.4221084 0.4342773 0.6640871
#> B D -0.3921116 0.2060007 -1.9034481 0.0569821
#> C D -0.1668590 0.1238819 -1.3469207 0.1780058
#> ci_lower ci_upper
#> -0.5780638 1.11462940
#> -0.9765735 0.24140380
#> -1.2584341 0.13173582
#> -0.6440051 1.01062924
#> -0.7958655 0.01164232
#> -0.4096630 0.07594494
#>
#> Contribution Matrix (% information from direct comparisons):
#> A vs B A vs C A vs D B vs C B vs D C vs D
#> A vs B 0 0.16 2.14 0 97.69 0.01
#> A vs C 0 99.87 0.11 0 0.00 0.02
#> A vs D 0 6.76 92.80 0 0.00 0.43
#> B vs C 0 4.89 1.81 0 93.29 0.02
#> B vs D 0 0.00 0.00 0 100.00 0.00
#> C vs D 0 72.80 26.93 0 0.00 0.27
#>
#> Model Fixed Effects (SE fallback applied when needed):
#> term estimate std_error z_value p_value se_source
#> d_B 0.198383855 0.1589885 1.24778748 0.21210889 hessian
#> d_C 0.388405127 0.2723588 1.42607902 0.15384553 hessian
#> d_D 0.596100819 0.3143572 1.89625336 0.05792655 hessian
#> severity 0.122704339 2.2649490 0.05417532 0.95679548 hessian
#> followup_months -0.007964814 0.1426709 -0.05582649 0.95548003 hessian
#>
#> Fit Statistics:
#> model_type heterogeneity n_treatments n_edges estimator QE QE_df
#> <NA> <NA> 6 6 MLE 0.106132 7
#> QE_p logLik Dev AIC BIC AICc
#> 0.9999972 3.609517 16.78097 6.780967 10.17531 -133.219
#>
#> Node-Splitting Assessment:
#> Q_node_split df p_value
#> 9.934809 6 0.1274243
#>
#> treatment_1 treatment_2 direct indirect direct_indirect_diff
#> A B 0.233333333 -0.034949478 0.2682828
#> A C 0.010410142 0.377994985 -0.3675848
#> A D 0.016375839 0.579724980 -0.5633491
#> B C 0.186666667 0.003354605 0.1833121
#> B D 0.002802677 0.394914287 -0.3921116
#> C D 0.020418327 0.187277366 -0.1668590
#> diff_se z p_value ci_lower ci_upper
#> 0.4318174 0.6212876 0.5344104 -0.5780638 1.11462940
#> 0.3107142 -1.1830320 0.2367965 -0.9765735 0.24140380
#> 0.3546417 -1.5885022 0.1121728 -1.2584341 0.13173582
#> 0.4221084 0.4342773 0.6640871 -0.6440051 1.01062924
#> 0.2060007 -1.9034481 0.0569821 -0.7958655 0.01164232
#> 0.1238819 -1.3469207 0.1780058 -0.4096630 0.07594494
#>
#> Treatment Ranking:
#> treatment mean_effect std_error mean_rank median_rank sucra sucr_a
#> D 0.5961008 0.3545198 1.10575 1 96.47500 96.47500
#> C 0.3884051 0.3106627 2.17850 2 60.71667 60.71667
#> B 0.1983839 0.1938489 3.01625 3 32.79167 32.79167
#> A 0.0000000 0.0000000 3.69950 4 10.01667 10.01667
#> p_best
#> 0.92000
#> 0.03575
#> 0.00625
#> 0.03800
#>
#> Contribution Decomposition (top rows):
#> Fixed-Effect Design Diagnostics:
#> $rank
#> [1] 3
#>
#> $ncol
#> [1] 3
#>
#> $condition_number
#> [1] 43.11314
#>
#> $info_condition_number
#> [1] 1.999933
#>
#> $singular
#> [1] FALSE
#>
#> $near_singular
#> [1] FALSE
#>
#> $recommendation
#> [1] "No major fixed-effect rank/conditioning issues detected."
#>
#>
#> Treatment Order:
#> rank treatment effect_vs_reference
#> 1 D 0.5961
#> 2 C 0.3884
#> 3 B 0.1984
#> 4 A 0.0000
#>
#> Direct / Indirect / Total Effects:
#> treatment_1 treatment_2 direct direct_se direct_n_studies direct_n_effects
#> A B 0.2333 0.2728 3 3
#> A C 0.0104 0.0040 2 2
#> A D 0.0164 0.0066 1 1
#> B C 0.1867 0.2749 2 2
#> B D 0.0028 0.0011 2 2
#> C D 0.0204 0.0157 2 2
#> direct_observed direct_observed_se indirect indirect_se indirect_n_paths
#> 0.2100 0.1095 -0.0349 0.3347 2
#> 0.3878 0.1491 0.3780 0.3107 2
#> 0.6100 0.2449 0.5797 0.3546 2
#> 0.1950 0.1414 0.0034 0.3203 2
#> 0.4082 0.1651 0.3949 0.2060 2
#> 0.2050 0.1581 0.1873 0.1229 2
#> indirect_n_studies indirect_n_effects total total_se total_n_studies
#> 7 7 0.1984 0.1938 9
#> 8 8 0.3884 0.3107 9
#> 8 9 0.5961 0.3545 9
#> 8 9 0.1900 0.1645 10
#> 8 8 0.3977 0.2060 10
#> 7 7 0.2077 0.1219 9
#> total_n_effects direct_ci_lower direct_ci_upper direct_observed_ci_lower
#> 10 -0.2155 0.6821 0.0298
#> 10 0.0038 0.0170 0.1426
#> 10 0.0056 0.0272 0.2071
#> 11 -0.2655 0.6388 -0.0376
#> 10 0.0009 0.0047 0.1365
#> 9 -0.0055 0.0463 -0.0551
#> direct_observed_ci_upper indirect_ci_lower indirect_ci_upper total_ci_lower
#> 0.3902 -0.5855 0.5156 -0.1205
#> 0.6330 -0.1330 0.8890 -0.1226
#> 1.0129 -0.0035 1.1630 0.0130
#> 0.4276 -0.5236 0.5303 -0.0806
#> 0.6798 0.0561 0.7338 0.0589
#> 0.4651 -0.0148 0.3894 0.0072
#> total_ci_upper
#> 0.5172
#> 0.8994
#> 1.1792
#> 0.4606
#> 0.7365
#> 0.4081
#>
#> Indirect SE note:
#> Indirect SEs are approximate; for mixed direct+indirect comparisons, var(indirect) is computed as var(total) + var(direct) assuming zero covariance.
#>
#> Heterogeneity Summary:
#> Overall Q:
#> $value
#> [,1]
#> [1,] 0.106132
#>
#> $df
#> [1] 7
#>
#> $p
#> [,1]
#> [1,] 0.9999972
#>
#>
#> Q by random effect level:
#> level Q df p_value n_groups
#> study_id 0.001859865 1 0.965601 1
#> component_id 0.000000000 0 NA 0
#>
#> Level-specific I2 / Tau2:
#> component tau2 Q df p_value n_groups I2
#> component_id 1e-06 0.000000000 0 NA 0 0.003080906
#> study_id 1e-06 0.001859865 1 0.965601 1 0.003080906
#>
#> Total I^2:
#> [1] 0.0008802783
#>
#> Within I^2:
#> [1] 0.003080906 0.003080906
#>
#> Between-study variance (tau^2):
#> component tau2
#> study_id 1e-06
#> component_id 1e-06The moderator specification is stored in the fitted object:
nma_ml$moderators
#> [1] "severity" "followup_months"League table of total effects:
nma_ml$league_table
#> A B C D
#> A 0.0000000 0.1983839 0.3884051 0.5961008
#> B -0.1983839 0.0000000 0.1900213 0.3977170
#> C -0.3884051 -0.1900213 0.0000000 0.2076957
#> D -0.5961008 -0.3977170 -0.2076957 0.0000000Formatted league table with confidence intervals:
network_league_table(nma_ml, digits = 2)
#> A B C
#> A "-" "0.20 [-0.12, 0.52]" "0.39 [-0.12, 0.90]"
#> B "-0.20 [-0.52, 0.12]" "-" "0.19 [-0.08, 0.46]"
#> C "-0.39 [-0.90, 0.12]" "-0.19 [-0.46, 0.08]" "-"
#> D "-0.60 [-1.18, -0.01]" "-0.40 [-0.74, -0.06]" "-0.21 [-0.41, -0.01]"
#> D
#> A "0.60 [0.01, 1.18]"
#> B "0.40 [0.06, 0.74]"
#> C "0.21 [0.01, 0.41]"
#> D "-"Per-comparison study contributions and indirect SEs:
nma_ml$effects[, c(
"treatment_1", "treatment_2",
"direct_n_studies", "indirect_n_studies", "total_n_studies",
"indirect", "indirect_se", "indirect_ci_lower", "indirect_ci_upper",
"total", "total_se", "total_ci_lower", "total_ci_upper"
)]
#> treatment_1 treatment_2 direct_n_studies indirect_n_studies total_n_studies
#> 1 A B 3 7 9
#> 2 A C 2 8 9
#> 3 A D 1 8 9
#> 4 B C 2 8 10
#> 5 B D 2 8 10
#> 6 C D 2 7 9
#> indirect indirect_se indirect_ci_lower indirect_ci_upper total
#> 1 -0.034949478 0.3346966 -0.585476459 0.5155775 0.1983839
#> 2 0.377994985 0.3106884 -0.133042007 0.8890320 0.3884051
#> 3 0.579724980 0.3545807 -0.003508424 1.1629584 0.5961008
#> 4 0.003354605 0.3203434 -0.523563438 0.5302726 0.1900213
#> 5 0.394914287 0.2059976 0.056078441 0.7337501 0.3977170
#> 6 0.187277366 0.1228768 -0.014836951 0.3893917 0.2076957
#> total_se total_ci_lower total_ci_upper
#> 1 0.1938489 -0.120469210 0.5172369
#> 2 0.3106627 -0.122589469 0.8993997
#> 3 0.3545198 0.012967718 1.1792339
#> 4 0.1645125 -0.080577667 0.4606202
#> 5 0.2059944 0.058886251 0.7365477
#> 6 0.1218634 0.007248219 0.4081432For flexible multilevel models, tau^2 is reported by
random level/component:
nma_ml$heterogeneity_summary$tau2
#> component tau2
#> 1 study_id 1e-06
#> 2 component_id 1e-06For multilevel models, Q is also broken out by
random/nested level:
nma_ml$heterogeneity_summary$Q_by_level
#> level Q df p_value n_groups
#> 1 study_id 0.001859865 1 0.965601 1
#> 2 component_id 0.000000000 0 NA 0Incoherence-factor assessment for multilevel NMA:
nma_ml$incoherence_assessment$global
#> Q_incoherence df p_value
#> 1 9.934809 6 0.1274243
nma_ml$incoherence_assessment$per_comparison
#> treatment_1 treatment_2 incoherence_factor if_se z p_value
#> 1 A B 0.2682828 0.4318174 0.6212876 0.5344104
#> 2 A C -0.3675848 0.3107142 -1.1830320 0.2367965
#> 3 A D -0.5633491 0.3546417 -1.5885022 0.1121728
#> 4 B C 0.1833121 0.4221084 0.4342773 0.6640871
#> 5 B D -0.3921116 0.2060007 -1.9034481 0.0569821
#> 6 C D -0.1668590 0.1238819 -1.3469207 0.1780058
#> ci_lower ci_upper
#> 1 -0.5780638 1.11462940
#> 2 -0.9765735 0.24140380
#> 3 -1.2584341 0.13173582
#> 4 -0.6440051 1.01062924
#> 5 -0.7958655 0.01164232
#> 6 -0.4096630 0.07594494Node-splitting for key direct/indirect contrasts:
nma_ml$node_splitting$global
#> Q_node_split df p_value
#> 1 9.934809 6 0.1274243
head(nma_ml$node_splitting$per_comparison)
#> treatment_1 treatment_2 direct indirect direct_indirect_diff
#> 1 A B 0.233333333 -0.034949478 0.2682828
#> 2 A C 0.010410142 0.377994985 -0.3675848
#> 3 A D 0.016375839 0.579724980 -0.5633491
#> 4 B C 0.186666667 0.003354605 0.1833121
#> 5 B D 0.002802677 0.394914287 -0.3921116
#> 6 C D 0.020418327 0.187277366 -0.1668590
#> diff_se z p_value ci_lower ci_upper
#> 1 0.4318174 0.6212876 0.5344104 -0.5780638 1.11462940
#> 2 0.3107142 -1.1830320 0.2367965 -0.9765735 0.24140380
#> 3 0.3546417 -1.5885022 0.1121728 -1.2584341 0.13173582
#> 4 0.4221084 0.4342773 0.6640871 -0.6440051 1.01062924
#> 5 0.2060007 -1.9034481 0.0569821 -0.7958655 0.01164232
#> 6 0.1238819 -1.3469207 0.1780058 -0.4096630 0.07594494Design-level inconsistency diagnostics:
design_inc <- network_design_inconsistency(nma_ml)
design_inc$Q_decomp
#> component Q df p_value
#> 1 whole_network 0.13360084 9 0.9999999
#> 2 within_designs 0.09222403 4 0.9989690
#> 3 between_designs 0.04137681 5 0.9999817
design_inc$by_design
#> design n_studies n_effects Q_within df p_value
#> A + B A + B 2 2 0.05142857 1 0.8205958
#> A + B + C A + B + C 1 2 0.00000000 0 NA
#> A + C A + C 1 1 0.00000000 0 NA
#> A + D A + D 1 1 0.00000000 0 NA
#> B + C B + C 2 2 0.00125000 1 0.9717964
#> B + D B + D 2 2 0.01454545 1 0.9040043
#> C + D C + D 2 2 0.02500000 1 0.8743671
design_inc$detach
#> detached_design Q_without_design df p_value delta_Q_between
#> 1 A + B + C 0.11259262 7 0.9999965 0.021008214
#> 2 A + B 0.07903771 7 0.9999990 0.054563125
#> 3 B + C 0.13111145 7 0.9999941 0.002489387
#> 4 A + D 0.13038105 8 0.9999993 0.003219787
#> 5 B + D 0.11503990 7 0.9999962 0.018560938
#> 6 C + D 0.10831017 7 0.9999970 0.025290670
#> 7 A + C 0.12513212 8 0.9999994 0.008468718
design_inc$heat_matrix
#> A vs B A vs C B vs C A vs D B vs D C vs D
#> A + B + C 0.01168144 0.009326771 0.000000000 0.000000000 0.00000000 0.00000000
#> A + B 0.05456313 0.000000000 0.000000000 0.000000000 0.00000000 0.00000000
#> B + C 0.00000000 0.000000000 0.002489387 0.000000000 0.00000000 0.00000000
#> A + D 0.00000000 0.000000000 0.000000000 0.003219787 0.00000000 0.00000000
#> B + D 0.00000000 0.000000000 0.000000000 0.000000000 0.01856094 0.00000000
#> C + D 0.00000000 0.000000000 0.000000000 0.000000000 0.00000000 0.02529067
#> A + C 0.00000000 0.008468718 0.000000000 0.000000000 0.00000000 0.00000000Contribution decomposition by study/source block:
head(nma_ml$contribution_decomposition$by_study)
#> [1] treatment_1 treatment_2 source_edge study
#> [5] effect_percentage
#> <0 rows> (or 0-length row.names)
head(nma_ml$contribution_decomposition$by_edge_block)
#> source_edge treatment_1 treatment_2 effect_percentage n_studies n_effects
#> 1 A vs B A B 4.499323e-10 3 3
#> 2 A vs B B C 3.807219e-10 3 3
#> 3 A vs B A C 2.202187e-11 3 3
#> 4 A vs B B D 5.142455e-13 3 3
#> 5 A vs B C D 5.277409e-09 3 3
#> 6 A vs B A D 1.818523e-08 3 3
#> sum_inverse_variance effective_information weighted_effective_information
#> 1 83.33333 2.941176 1.323330e-11
#> 2 83.33333 2.941176 1.119770e-11
#> 3 83.33333 2.941176 6.477021e-13
#> 4 83.33333 2.941176 1.512487e-14
#> 5 83.33333 2.941176 1.552179e-10
#> 6 83.33333 2.941176 5.348598e-10
head(nma_ml$contribution_decomposition$effective_information$by_edge)
#> source_edge n_studies n_effects sum_inverse_variance effective_information
#> 1 A vs B 3 3 83.33333 2.941176
#> 2 A vs C 2 2 45.00000 1.975610
#> 3 A vs D 1 1 16.66667 1.000000
#> 4 B vs C 2 2 50.00000 2.000000
#> 5 B vs D 2 2 36.66667 1.983607
#> 6 C vs D 2 2 40.00000 2.000000
head(nma_ml$contribution_decomposition$effective_information$by_edge_study)
#> source_edge study n_effects sum_inverse_variance effective_information
#> 1 A vs B s1 1 25.00000 1
#> 2 A vs B s10 1 25.00000 1
#> 3 A vs B s2 1 33.33333 1
#> 4 A vs C s1 1 20.00000 1
#> 5 A vs C s7 1 25.00000 1
#> 6 A vs D s4 1 16.66667 1
head(nma_ml$contribution_decomposition$effective_information$by_comparison)
#> treatment_1 treatment_2 weighted_effective_information
#> 1 A B NA
#> 2 B C NA
#> 3 A C NA
#> 4 B D NA
#> 5 C D NA
#> 6 A D NAFit statistics:
nma_ml$fit_stats
#> model_type heterogeneity n_treatments n_edges estimator QE QE_df
#> 1 <NA> <NA> 6 6 MLE 0.106132 7
#> QE_p logLik Dev AIC BIC AICc
#> 1 0.9999972 3.609517 16.78097 6.780967 10.17531 -133.219Ranking output:
nma_ml$ranking$rankings
#> treatment mean_effect std_error mean_rank median_rank sucra sucr_a
#> D D 0.5961008 0.3545198 1.10575 1 96.47500 96.47500
#> C C 0.3884051 0.3106627 2.17850 2 60.71667 60.71667
#> B B 0.1983839 0.1938489 3.01625 3 32.79167 32.79167
#> A A 0.0000000 0.0000000 3.69950 4 10.01667 10.01667
#> p_best
#> D 0.92000
#> C 0.03575
#> B 0.00625
#> A 0.03800
nma_ml$ranking$rank_probability[1:4, , drop = FALSE]
#> rank_1 rank_2 rank_3 rank_4
#> A 0.03800 0.05300 0.08050 0.82850
#> B 0.00625 0.07175 0.82150 0.10050
#> C 0.03575 0.81625 0.08175 0.06625
#> D 0.92000 0.05900 0.01625 0.00475
nma_ml$league_table_ordered
#> treatment_1 treatment_2 total total_se total_ci_lower total_ci_upper
#> 3 A D 0.5961008 0.3545198 0.012967718 1.1792339
#> 5 B D 0.3977170 0.2059944 0.058886251 0.7365477
#> 2 A C 0.3884051 0.3106627 -0.122589469 0.8993997
#> 6 C D 0.2076957 0.1218634 0.007248219 0.4081432
#> 1 A B 0.1983839 0.1938489 -0.120469210 0.5172369
#> 4 B C 0.1900213 0.1645125 -0.080577667 0.4606202
head(nma_ml$league_table_se)
#> A B C D
#> A 0.0000000 0.1938489 0.3106627 0.3545198
#> B 0.1938489 0.0000000 0.1645125 0.2059944
#> C 0.3106627 0.1645125 0.0000000 0.1218634
#> D 0.3545198 0.2059944 0.1218634 0.0000000Prediction intervals for all total network effects:
network_prediction_intervals(nma_ml, level = 0.90)
#> treatment_1 treatment_2 total total_se tau2_prediction prediction_se
#> 1 A B 0.1983839 0.1938489 1e-06 0.1938515
#> 2 A C 0.3884051 0.3106627 1e-06 0.3106643
#> 3 A D 0.5961008 0.3545198 1e-06 0.3545212
#> 4 B C 0.1900213 0.1645125 1e-06 0.1645155
#> 5 B D 0.3977170 0.2059944 1e-06 0.2059969
#> 6 C D 0.2076957 0.1218634 1e-06 0.1218675
#> prediction_lower prediction_upper
#> 1 -0.12047345 0.5172412
#> 2 -0.12259212 0.8994024
#> 3 0.01296540 1.1792362
#> 4 -0.08058267 0.4606252
#> 5 0.05888226 0.7365517
#> 6 0.00724147 0.4081499Prediction intervals are also available through
predict() for selected contrasts:
predict(
nma_ml,
newdata = data.frame(
treatment_1 = c("A", "B"),
treatment_2 = c("D", "C"),
severity = c(0.5, 0.5),
followup_months = c(6, 6)
),
se.fit = TRUE,
interval = "prediction",
level = 0.90
)
#> treatment_1 treatment_2 estimate se prediction_se tau2_prediction
#> 1 A D 0.6096641 0.1341185 0.1341222 1e-06
#> 2 B C 0.2035846 0.2254526 0.2254548 1e-06
#> lower upper
#> 1 0.3890527 0.8302756
#> 2 -0.1672556 0.5744247Level-specific heterogeneity:
nma_ml$heterogeneity_summary$level_heterogeneity
#> component tau2 Q df p_value n_groups I2
#> 1 component_id 1e-06 0.000000000 0 NA 0 0.003080906
#> 2 study_id 1e-06 0.001859865 1 0.965601 1 0.003080906Contribution matrix for multilevel NMA:
nma_ml$contribution_matrix
#> A vs B A vs C A vs D B vs C B vs D
#> A vs B 4.499323e-10 1.563275e-01 2.144864e+00 3.812357e-10 9.768878e+01
#> A vs C 2.202187e-11 9.986814e+01 1.123532e-01 6.890900e-10 2.739690e-05
#> A vs D 1.818523e-08 6.762164e+00 9.280012e+01 1.534226e-08 3.903011e-03
#> B vs C 3.807219e-10 4.885075e+00 1.807110e+00 6.710233e-10 9.328982e+01
#> B vs D 5.142455e-13 1.023784e-09 2.423296e-09 4.917517e-13 1.000000e+02
#> C vs D 5.277409e-09 7.279891e+01 2.693127e+01 9.482879e-09 1.688052e-03
#> C vs D
#> A vs B 1.002643e-02
#> A vs C 1.948361e-02
#> A vs D 4.338121e-01
#> B vs C 1.799203e-02
#> B vs D 1.688253e-11
#> C vs D 2.681312e-01Multilevel Variants
nested_levels can be used to add deeper random-effect
nesting. The template below is shown without evaluation so the vignette
documents the API without paying for another full fit during
compilation.
nma_dat$site <- rep(c("north", "south", "west"), length.out = nrow(nma_dat))
nma_dat$cohort <- rep(c("c1", "c2"), length.out = nrow(nma_dat))
nma_ml_nested <- network_meta(
data = nma_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multilevel",
tau_components = "comparison",
moderators = ~ severity + followup_months,
nested_levels = c("site", "cohort"),
estimation_method = "MLE"
)
summary(nma_ml_nested)You can also express outcome-within-study nesting explicitly:
nma_nested_dat <- nma_dat
nma_nested_dat$outcome <- rep(c("pain", "function"), length.out = nrow(nma_nested_dat))
nma_nested_dat$comparison_within_outcome <- ave(
seq_len(nrow(nma_nested_dat)),
nma_nested_dat$study,
nma_nested_dat$outcome,
FUN = seq_along
)
nma_ml_outcome_nested <- network_meta(
data = nma_nested_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multilevel",
tau_components = "comparison",
nested_levels = c("outcome", "comparison_within_outcome"),
estimation_method = "MLE"
)
nma_ml_outcome_nested$fit$formula
nma_ml_outcome_nested$heterogeneity_summary$Q_by_level
nma_ml_outcome_nested$heterogeneity_summary$tau2Multilevel and robustID
In network_meta(), robustID is restricted
to model_type = "multivariate". For multilevel models,
clustering is handled through random effects (study_id,
tau_components, nested_levels), so
robustID now returns a clear error.
Forest Plots for Network Effects
network_graph_plot() draws the direct treatment network
using base R. Edge widths and node sizes can reflect the number of
direct studies or direct effect sizes.
network_graph_plot(
nma_ml,
node_size = "studies",
edge_width = "studies",
edge_label = TRUE,
main = "Treatment Network"
)
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1
#> Warning in text.default(mean(c(nodes$x[node_index[ii]], nodes$x[node_index_2[ii]])), : length(labels) > max(length(x), length(y));
#> 'labels' truncated to length 1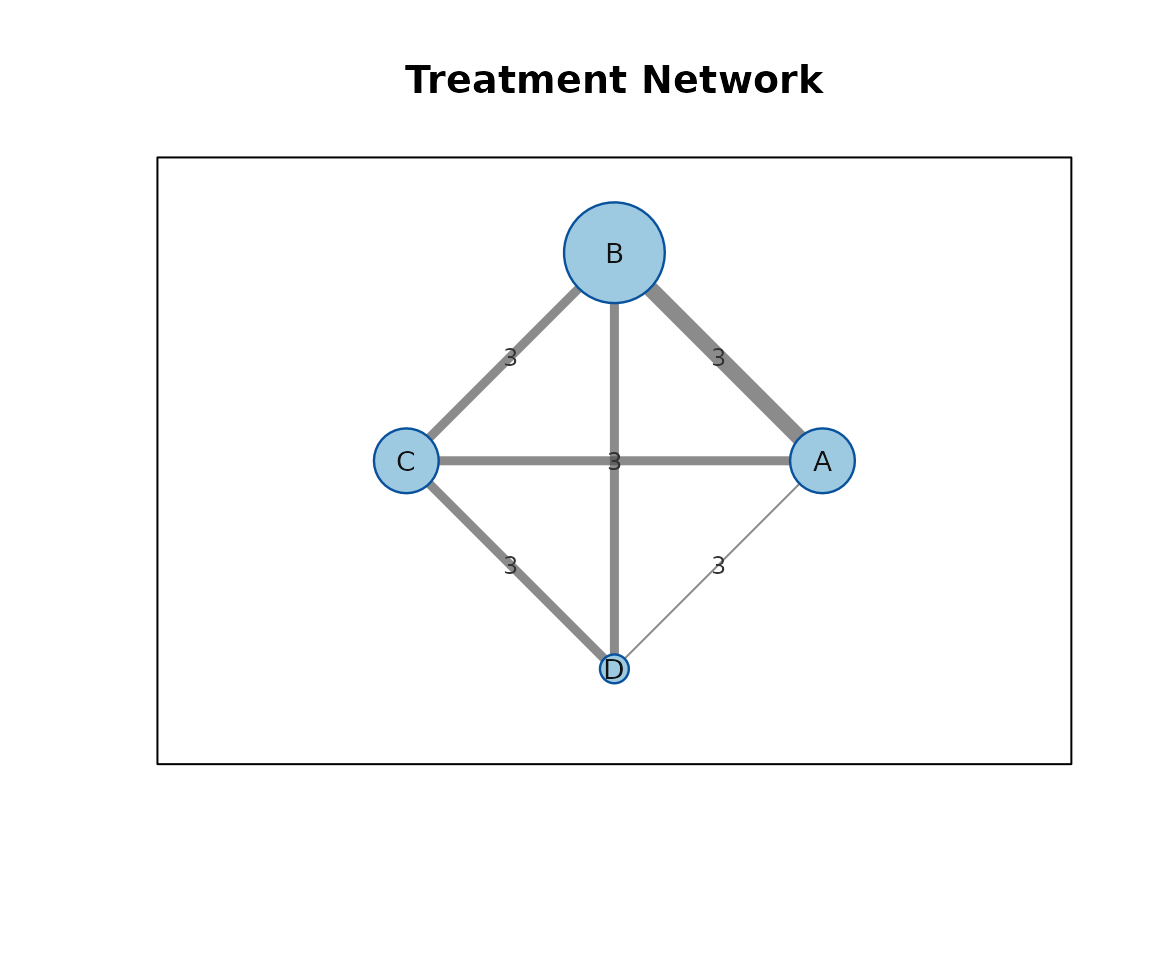
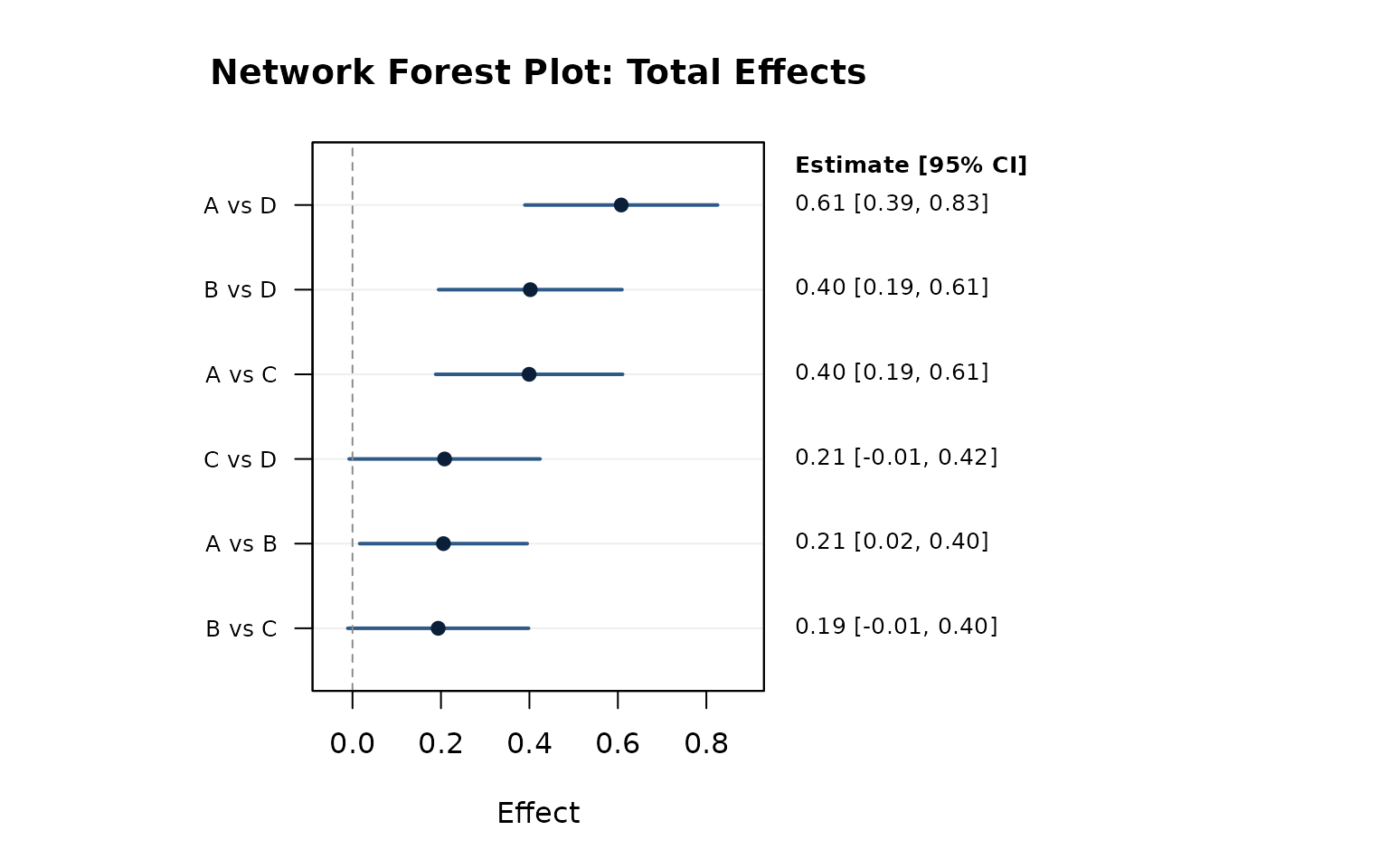
network_forest_plot() provides a standard forest plot
for one selected effect type ("direct",
"indirect", or "total").
network_forest_plot(
nma_ml,
effect_type = "total",
order_by = "effect",
right_digits = 2,
main = "Network Forest Plot: Total Effects"
)
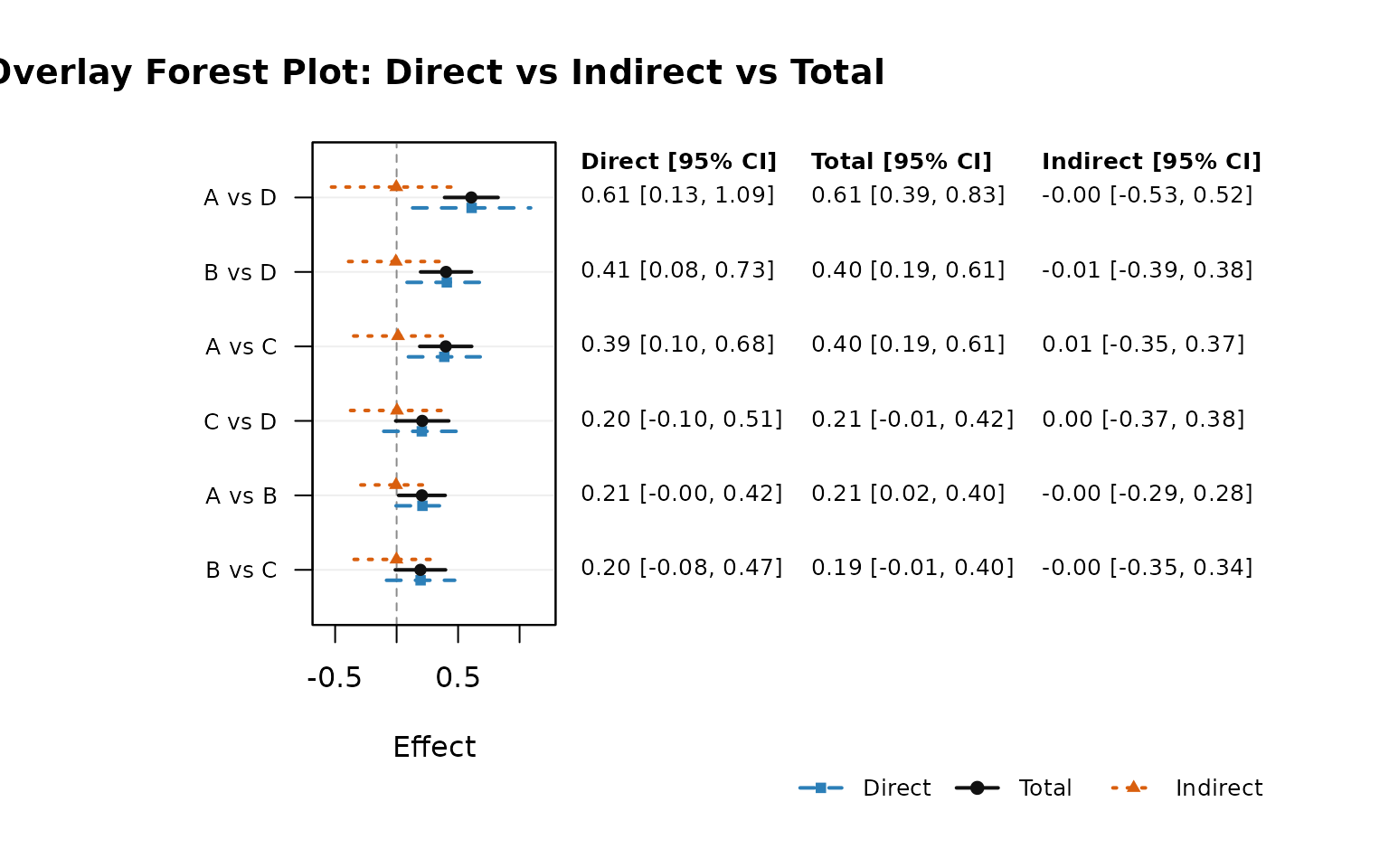
network_forest_overlay_plot() draws direct, total, and
indirect effects for each comparison using nearly overlapping lines for
fast visual comparison.
network_forest_overlay_plot(
nma_ml,
order_by = "total",
right_label = "all",
right_digits = 2,
main = "Overlay Forest Plot: Direct vs Indirect vs Total"
)
Contribution Heatmap (Base R)
network_contribution_heatmap() visualizes the percentage
information contributed by each direct comparison estimate to each
network comparison.
network_contribution_heatmap(
nma_ml,
main = "Contribution Matrix Heatmap",
show_values = FALSE
)
Arm-Level Pairwise Conversion
network_pairwise_data() converts arm-level binary or
continuous data to the contrast-level format used by
network_meta(). Binary examples can use log odds ratios
("OR"), log risk ratios ("RR"), or risk
differences ("RD"). Continuous examples can use mean
differences ("MD").
arm_dat <- data.frame(
study = c("s1", "s1", "s1", "s2", "s2"),
treatment = c("A", "B", "C", "A", "B"),
event = c(4, 8, 10, 3, 6),
n = c(40, 42, 41, 35, 37)
)
network_pairwise_data(
arm_dat,
study_id = "study",
treatment = "treatment",
event = "event",
n = "n",
effect_measure = "OR"
)
#> study treatment_1 treatment_2 effect variance effect_measure
#> 1 s1 A B 0.7503056 0.4321895 OR
#> 2 s1 A C 1.0658225 0.4100358 OR
#> 3 s1 B C 0.3155169 0.2866698 OR
#> 4 s2 A B 0.7248959 0.5635081 ORComparison-Adjusted Funnel Plot
Small-study diagnostics for network meta-analysis can center each observed contrast around the fitted network estimate for its comparison.
network_funnel_plot(nma_ml)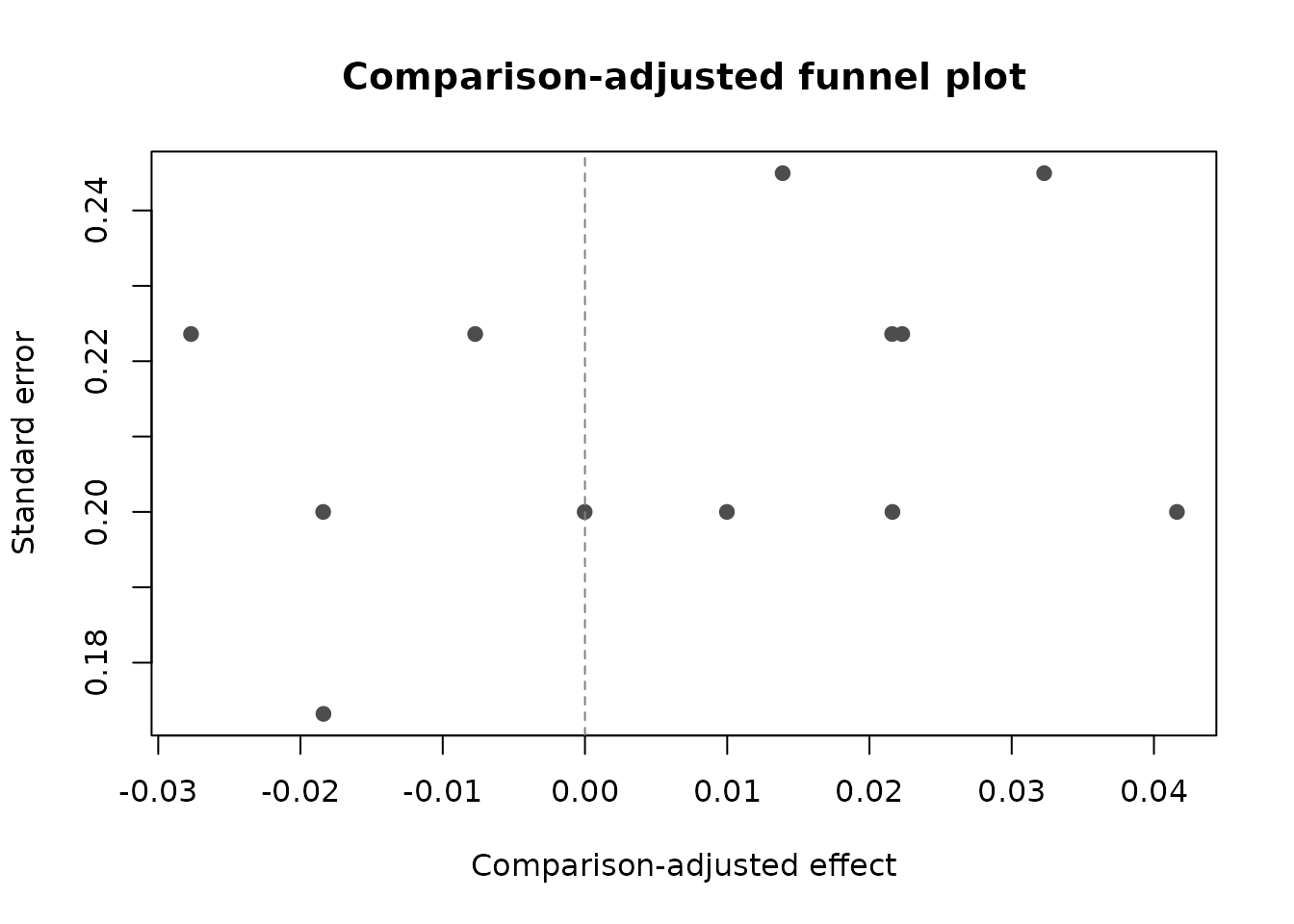
network_bias_test(nma_ml)$intercept
#> term estimate std_error t_value p_value
#> 1 (Intercept) 0.3492353 0.3109481 1.123131 0.2876223Component Network Meta-analysis
When treatment names represent combinations of components,
network_component_meta() fits an additive component model
from a fitted nma_mars object. The example below treats
placebo as inactive and estimates the separate contributions of
components A and B.
component_dat <- data.frame(
study = paste0("s", 1:8),
trt1 = c("Placebo", "Placebo", "Placebo", "A", "B", "Placebo", "A", "B"),
trt2 = c("A", "B", "A+B", "A+B", "A+B", "A+B", "A+B", "A+B"),
yi = c(0.20, 0.30, 0.52, 0.31, 0.19, 0.48, 0.29, 0.21),
vi = rep(0.04, 8)
)
component_nma <- network_meta(
data = component_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "Placebo",
model_type = "wls"
)
component_fit <- network_component_meta(component_nma, inactive = "Placebo")
component_fit$component_effects
#> component estimate std_error ci_lower ci_upper z_value p_value
#> A A 0.2 0.09759001 0.0087271 0.3912729 2.049390 0.040423979
#> B B 0.3 0.09759001 0.1087271 0.4912729 3.074085 0.002111491
component_fit$combinations
#> treatment estimate std_error ci_lower ci_upper observed_nma_effect
#> A A 0.2 0.09759001 0.0087271 0.3912729 0.2
#> A+B A+B 0.5 0.10690450 0.2904710 0.7095290 0.5
#> B B 0.3 0.09759001 0.1087271 0.4912729 0.3
#> Placebo Placebo 0.0 0.00000000 0.0000000 0.0000000 0.0
#> residual
#> A 0.000000e+00
#> A+B -1.110223e-16
#> B 0.000000e+00
#> Placebo 0.000000e+00
component_fit$fit_stats
#> Q_additive df p_value rank singular
#> 1 1.848893e-30 1 1 2 FALSEnetwork_complex_effect() estimates arbitrary complex
interventions or comparisons from the component model.
network_complex_effect(component_fit, "A + B")
#> intervention comparator estimate std_error ci_lower ci_upper
#> 1 A + B Placebo 0.5 0.1069045 0.290471 0.709529
network_complex_effect(component_fit, intervention = "A+B", comparator = "A")
#> intervention comparator estimate std_error ci_lower ci_upper
#> 1 A+B A 0.3 0.09759001 0.1087271 0.4912729
network_complex_effect(component_fit, 2)
#> intervention comparator estimate std_error ci_lower ci_upper
#> 1 A + B Placebo 0.5 0.1069045 0.290471 0.709529Speed Controls
For larger networks, control can enable sparse matrix
algebra, reuse cached intermediate matrices, and parallelize selected
post-fit computations.
nma_ml_fast <- network_meta(
data = nma_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multilevel",
tau_components = "comparison",
moderators = ~ severity + followup_months,
estimation_method = "MLE",
control = list(
use_sparse = TRUE,
reuse_inverses = TRUE,
parallel = 2
)
)
nma_ml_fast$controlMultivariate Network Meta-analysis
You can also fit in multivariate mode using
model_type = "multivariate". Use
within_varcov_type to choose the within-study covariance
structure used by the multivariate engine (for example
"multilevel"/"univariate" or metric-based
options such as "log_or" or
"smd_shared_control" when those required fields are
available). This example also includes moderators so one fitted object
shows the full cycle.
nma_mv <- suppressWarnings(network_meta(
data = nma_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multivariate",
heterogeneity = "flexible",
moderators = c("severity", "followup_months"),
estimation_method = "MLE",
within_varcov_type = "multilevel"
))
summary(nma_mv)
#> Network Meta-analysis (MARS)
#> Model Type: multivariate
#> Heterogeneity: flexible
#> Moderators: severity, followup_months
#> Within-study covariance type: multilevel
#> Stored within-study covariance blocks: 11
#> Reference: A
#> Treatments: A, B, C, D
#>
#> Evidence Summary:
#> $n_comparisons_total
#> [1] 6
#>
#> $n_comparisons_with_direct
#> [1] 6
#>
#> $n_comparisons_with_indirect
#> [1] 6
#>
#> $n_comparisons_indirect_only
#> [1] 0
#>
#> $n_direct_effects
#> [1] 12
#>
#> $n_indirect_effects
#> [1] 48
#>
#>
#> Incoherence-Factor Assessment:
#> Global test:
#> Q_incoherence df p_value
#> 9.935036 6 0.1274146
#>
#> Per-comparison incoherence factors:
#> treatment_1 treatment_2 incoherence_factor if_se z p_value
#> A B 0.2682826 0.4318164 0.6212886 0.53440973
#> A C -0.3675853 0.3107103 -1.1830483 0.23679000
#> A D -0.5633495 0.3546375 -1.5885219 0.11216838
#> B C 0.1833119 0.4221076 0.4342776 0.66408688
#> B D -0.3921117 0.2059983 -1.9034708 0.05697914
#> C D -0.1668590 0.1238805 -1.3469346 0.17800131
#> ci_lower ci_upper
#> -0.5780619 1.11462711
#> -0.9765662 0.24139567
#> -1.2584263 0.13172731
#> -0.6440038 1.01062752
#> -0.7958610 0.01163749
#> -0.4096603 0.07594241
#>
#> Contribution Matrix (% information from direct comparisons):
#> A vs B A vs C A vs D B vs C B vs D C vs D
#> A vs B 0 0.16 2.14 0 97.69 0.01
#> A vs C 0 99.87 0.11 0 0.00 0.02
#> A vs D 0 6.76 92.80 0 0.00 0.43
#> B vs C 0 4.89 1.81 0 93.29 0.02
#> B vs D 0 0.00 0.00 0 100.00 0.00
#> C vs D 0 72.80 26.93 0 0.00 0.27
#>
#> Model Fixed Effects (SE fallback applied when needed):
#> term estimate std_error z_value p_value
#> factor(data[[effectID]])A vs B 0.21372546 0.2217672 0.9637377 0.3351774
#> factor(data[[effectID]])A vs C 0.36372546 0.3658167 0.9942833 0.3200849
#> factor(data[[effectID]])A vs D 0.58372546 0.3724922 1.5670808 0.1170958
#> factor(data[[effectID]])B vs C 0.20725491 0.2187216 0.9475742 0.3433463
#> factor(data[[effectID]])B vs D 0.42973857 0.3122505 1.3762621 0.1687405
#> factor(data[[effectID]])C vs D 0.20549028 0.1686609 1.2183633 0.2230860
#> severity 0.37254908 2.8438590 0.1310012 0.8957743
#> followup_months -0.02666667 0.1882336 -0.1416679 0.8873423
#> se_source
#> hessian
#> hessian
#> hessian
#> hessian
#> hessian
#> hessian
#> hessian
#> hessian
#>
#> Fit Statistics:
#> model_type heterogeneity n_treatments n_edges estimator QE QE_df
#> <NA> <NA> 9 6 MLE 9.140389 6
#> QE_p logLik Dev AIC BIC AICc
#> 0.1658353 7.234778 9.530444 13.53044 20.31914 -126.4696
#>
#> Node-Splitting Assessment:
#> Q_node_split df p_value
#> 9.935036 6 0.1274146
#>
#> treatment_1 treatment_2 direct indirect direct_indirect_diff
#> A B 0.233333333 -0.034949264 0.2682826
#> A C 0.010410142 0.377995400 -0.3675853
#> A D 0.016375839 0.579725324 -0.5633495
#> B C 0.186666667 0.003354805 0.1833119
#> B D 0.002802677 0.394914417 -0.3921117
#> C D 0.020418327 0.187277295 -0.1668590
#> diff_se z p_value ci_lower ci_upper
#> 0.4318164 0.6212886 0.53440973 -0.5780619 1.11462711
#> 0.3107103 -1.1830483 0.23679000 -0.9765662 0.24139567
#> 0.3546375 -1.5885219 0.11216838 -1.2584263 0.13172731
#> 0.4221076 0.4342776 0.66408688 -0.6440038 1.01062752
#> 0.2059983 -1.9034708 0.05697914 -0.7958610 0.01163749
#> 0.1238805 -1.3469346 0.17800131 -0.4096603 0.07594241
#>
#> Treatment Ranking:
#> treatment mean_effect std_error mean_rank median_rank sucra sucr_a
#> D 0.5961012 0.3545156 1.123 1 95.90000 95.90000
#> C 0.3884055 0.3106587 2.170 2 61.00000 61.00000
#> B 0.1983841 0.1938465 3.008 3 33.06667 33.06667
#> A 0.0000000 0.0000000 3.699 4 10.03333 10.03333
#> p_best
#> 0.90775
#> 0.03975
#> 0.00825
#> 0.04425
#>
#> Contribution Decomposition (top rows):
#> Fixed-Effect Design Diagnostics:
#> $rank
#> [1] 3
#>
#> $ncol
#> [1] 3
#>
#> $condition_number
#> [1] 43.11297
#>
#> $info_condition_number
#> [1] 1.999967
#>
#> $singular
#> [1] FALSE
#>
#> $near_singular
#> [1] FALSE
#>
#> $recommendation
#> [1] "No major fixed-effect rank/conditioning issues detected."
#>
#>
#> Treatment Order:
#> rank treatment effect_vs_reference
#> 1 D 0.5961
#> 2 C 0.3884
#> 3 B 0.1984
#> 4 A 0.0000
#>
#> Direct / Indirect / Total Effects:
#> treatment_1 treatment_2 direct direct_se direct_n_studies direct_n_effects
#> A B 0.2333 0.2728 3 3
#> A C 0.0104 0.0040 2 2
#> A D 0.0164 0.0066 1 1
#> B C 0.1867 0.2749 2 2
#> B D 0.0028 0.0011 2 2
#> C D 0.0204 0.0157 2 2
#> direct_observed direct_observed_se indirect indirect_se indirect_n_paths
#> 0.2100 0.1095 -0.0349 0.3347 2
#> 0.3878 0.1491 0.3780 0.3107 2
#> 0.6100 0.2449 0.5797 0.3546 2
#> 0.1950 0.1414 0.0034 0.3203 2
#> 0.4082 0.1651 0.3949 0.2060 2
#> 0.2050 0.1581 0.1873 0.1229 2
#> indirect_n_studies indirect_n_effects total total_se total_n_studies
#> 7 7 0.1984 0.1938 9
#> 8 8 0.3884 0.3107 9
#> 8 9 0.5961 0.3545 9
#> 8 9 0.1900 0.1645 10
#> 8 8 0.3977 0.2060 10
#> 7 7 0.2077 0.1219 9
#> total_n_effects direct_ci_lower direct_ci_upper direct_observed_ci_lower
#> 10 -0.3014 0.7681 -0.0047
#> 10 0.0026 0.0183 0.0956
#> 10 0.0035 0.0293 0.1299
#> 11 -0.3521 0.7254 -0.0822
#> 10 0.0006 0.0050 0.0845
#> 9 -0.0104 0.0513 -0.1049
#> direct_observed_ci_upper indirect_ci_lower indirect_ci_upper total_ci_lower
#> 0.4247 -0.6909 0.6210 -0.1815
#> 0.6800 -0.2309 0.9869 -0.2205
#> 1.0901 -0.1152 1.2747 -0.0987
#> 0.4722 -0.6245 0.6312 -0.1324
#> 0.7319 -0.0088 0.7987 -0.0060
#> 0.5149 -0.0536 0.4281 -0.0311
#> total_ci_upper
#> 0.5783
#> 0.9973
#> 1.2909
#> 0.5125
#> 0.8015
#> 0.4465
#>
#> Indirect SE note:
#> Indirect SEs are approximate; for mixed direct+indirect comparisons, var(indirect) is computed as var(total) + var(direct) assuming zero covariance.
#>
#> Heterogeneity Summary:
#> Overall Q:
#> $value
#> [,1]
#> [1,] 9.140389
#>
#> $df
#> [1] 6
#>
#> $p
#> [,1]
#> [1,] 0.1658353
#>
#>
#> Within-level Q:
#> $`A vs B`
#> QE_value QE_df QE_p
#> 0.0400000 2.0000000 0.9801987
#>
#> $`A vs C`
#> QE_value QE_df QE_p
#> 0.0160000 1.0000000 0.8993432
#>
#> $`A vs D`
#> [1] "only one effect size in this category"
#>
#> $`B vs C`
#> QE_value QE_df QE_p
#> 0.001111111 1.000000000 0.973408772
#>
#> $`B vs D`
#> QE_value QE_df QE_p
#> 0.01777778 1.00000000 0.89392977
#>
#> $`C vs D`
#> QE_value QE_df QE_p
#> 0.0250000 1.0000000 0.8743671
#>
#>
#> Total I^2:
#> [1] 93.43997
#>
#> Within I^2:
#> [1] 0.002136528 0.002136528 95.528876009 95.528876009 95.528876009
#> [6] 95.528876009
#>
#> Between-study variance (tau^2):
#> component tau2
#> A vs B 1e-06
#> A vs C 1e-06
#> A vs D 1e+00
#> B vs C 1e+00
#> B vs D 1e+00
#> C vs D 1e+00The moderator specification is stored in the fitted object:
nma_mv$moderators
#> [1] "severity" "followup_months"The output structure is the same, including effects, heterogeneity summaries, and treatment order:
nma_mv$effects
#> treatment_1 treatment_2 direct direct_se direct_n_studies
#> 1 A B 0.233333333 0.272845092 3
#> 2 A C 0.010410142 0.004001911 2
#> 3 A D 0.016375839 0.006575811 1
#> 4 B C 0.186666667 0.274873708 2
#> 5 B D 0.002802677 0.001133923 2
#> 6 C D 0.020418327 0.015748395 2
#> direct_n_effects direct_observed direct_observed_se indirect indirect_se
#> 1 3 0.2100000 0.1095445 -0.034949264 0.3346953
#> 2 2 0.3877778 0.1490712 0.377995400 0.3106845
#> 3 1 0.6100000 0.2449490 0.579725324 0.3545766
#> 4 2 0.1950000 0.1414214 0.003354805 0.3203424
#> 5 2 0.4081818 0.1651446 0.394914417 0.2059952
#> 6 2 0.2050000 0.1581139 0.187277295 0.1228754
#> indirect_n_paths indirect_n_studies indirect_n_effects total total_se
#> 1 2 7 7 0.1983841 0.1938465
#> 2 2 8 8 0.3884055 0.3106587
#> 3 2 8 9 0.5961012 0.3545156
#> 4 2 8 9 0.1900215 0.1645105
#> 5 2 8 8 0.3977171 0.2059920
#> 6 2 7 7 0.2076956 0.1218621
#> total_n_studies total_n_effects direct_ci_lower direct_ci_upper
#> 1 9 10 -0.3014332211 0.768099888
#> 2 9 10 0.0025665395 0.018253744
#> 3 9 10 0.0034874854 0.029264192
#> 4 10 11 -0.3520759020 0.725409235
#> 5 10 10 0.0005802281 0.005025125
#> 6 9 9 -0.0104479598 0.051284613
#> direct_observed_ci_lower direct_observed_ci_upper indirect_ci_lower
#> 1 -0.004703297 0.4247033 -0.690939943
#> 2 0.095603598 0.6799520 -0.230935013
#> 3 0.129908832 1.0900912 -0.115231972
#> 4 -0.082180765 0.4721808 -0.624504774
#> 5 0.084504419 0.7318592 -0.008828698
#> 6 -0.104897516 0.5148975 -0.053554142
#> indirect_ci_upper total_ci_lower total_ci_upper
#> 1 0.6210414 -0.181548166 0.5783163
#> 2 0.9869258 -0.220474353 0.9972854
#> 3 1.2746826 -0.098736612 1.2909389
#> 4 0.6312144 -0.132413166 0.5124561
#> 5 0.7986575 -0.006019905 0.8014541
#> 6 0.4281087 -0.031149628 0.4465409
nma_mv$evidence_summary
#> $n_comparisons_total
#> [1] 6
#>
#> $n_comparisons_with_direct
#> [1] 6
#>
#> $n_comparisons_with_indirect
#> [1] 6
#>
#> $n_comparisons_indirect_only
#> [1] 0
#>
#> $n_direct_effects
#> [1] 12
#>
#> $n_indirect_effects
#> [1] 48
nma_mv$indirect_se_note
#> [1] "Indirect SEs are approximate; for mixed direct+indirect comparisons, var(indirect) is computed as var(total) + var(direct) assuming zero covariance."
nma_mv$heterogeneity_summary
#> $Q_total
#> $Q_total$value
#> [,1]
#> [1,] 9.140389
#>
#> $Q_total$df
#> [1] 6
#>
#> $Q_total$p
#> [,1]
#> [1,] 0.1658353
#>
#>
#> $Q_within
#> $Q_within$`A vs B`
#> QE_value QE_df QE_p
#> 0.0400000 2.0000000 0.9801987
#>
#> $Q_within$`A vs C`
#> QE_value QE_df QE_p
#> 0.0160000 1.0000000 0.8993432
#>
#> $Q_within$`A vs D`
#> [1] "only one effect size in this category"
#>
#> $Q_within$`B vs C`
#> QE_value QE_df QE_p
#> 0.001111111 1.000000000 0.973408772
#>
#> $Q_within$`B vs D`
#> QE_value QE_df QE_p
#> 0.01777778 1.00000000 0.89392977
#>
#> $Q_within$`C vs D`
#> QE_value QE_df QE_p
#> 0.0250000 1.0000000 0.8743671
#>
#>
#> $Q_by_level
#> NULL
#>
#> $I2_total
#> [1] 93.43997
#>
#> $I2_within
#> [1] 0.002136528 0.002136528 95.528876009 95.528876009 95.528876009
#> [6] 95.528876009
#>
#> $level_heterogeneity
#> NULL
#>
#> $tau2
#> component tau2
#> 1 A vs B 1e-06
#> 2 A vs C 1e-06
#> 3 A vs D 1e+00
#> 4 B vs C 1e+00
#> 5 B vs D 1e+00
#> 6 C vs D 1e+00
nma_mv$heterogeneity_summary$tau2
#> component tau2
#> 1 A vs B 1e-06
#> 2 A vs C 1e-06
#> 3 A vs D 1e+00
#> 4 B vs C 1e+00
#> 5 B vs D 1e+00
#> 6 C vs D 1e+00
nma_mv$incoherence_assessment$global
#> Q_incoherence df p_value
#> 1 9.935036 6 0.1274146
nma_mv$incoherence_assessment$per_comparison
#> treatment_1 treatment_2 incoherence_factor if_se z p_value
#> 1 A B 0.2682826 0.4318164 0.6212886 0.53440973
#> 2 A C -0.3675853 0.3107103 -1.1830483 0.23679000
#> 3 A D -0.5633495 0.3546375 -1.5885219 0.11216838
#> 4 B C 0.1833119 0.4221076 0.4342776 0.66408688
#> 5 B D -0.3921117 0.2059983 -1.9034708 0.05697914
#> 6 C D -0.1668590 0.1238805 -1.3469346 0.17800131
#> ci_lower ci_upper
#> 1 -0.5780619 1.11462711
#> 2 -0.9765662 0.24139567
#> 3 -1.2584263 0.13172731
#> 4 -0.6440038 1.01062752
#> 5 -0.7958610 0.01163749
#> 6 -0.4096603 0.07594241
nma_mv$contribution_matrix
#> A vs B A vs C A vs D B vs C B vs D
#> A vs B 4.499323e-10 1.563275e-01 2.144864e+00 3.812357e-10 9.768878e+01
#> A vs C 2.202187e-11 9.986814e+01 1.123532e-01 6.890900e-10 2.739690e-05
#> A vs D 1.818523e-08 6.762164e+00 9.280012e+01 1.534226e-08 3.903011e-03
#> B vs C 3.807219e-10 4.885075e+00 1.807110e+00 6.710233e-10 9.328982e+01
#> B vs D 5.142455e-13 1.023784e-09 2.423296e-09 4.917517e-13 1.000000e+02
#> C vs D 5.277409e-09 7.279891e+01 2.693127e+01 9.482879e-09 1.688052e-03
#> C vs D
#> A vs B 1.002643e-02
#> A vs C 1.948361e-02
#> A vs D 4.338121e-01
#> B vs C 1.799203e-02
#> B vs D 1.688253e-11
#> C vs D 2.681312e-01
nma_mv$treatment_order
#> rank treatment effect_vs_reference
#> 1 1 D 0.5961012
#> 2 2 C 0.3884055
#> 3 3 B 0.1983841
#> 4 4 A 0.0000000
nma_mv$node_splitting$global
#> Q_node_split df p_value
#> 1 9.935036 6 0.1274146
nma_mv$ranking$rankings
#> treatment mean_effect std_error mean_rank median_rank sucra sucr_a
#> D D 0.5961012 0.3545156 1.123 1 95.90000 95.90000
#> C C 0.3884055 0.3106587 2.170 2 61.00000 61.00000
#> B B 0.1983841 0.1938465 3.008 3 33.06667 33.06667
#> A A 0.0000000 0.0000000 3.699 4 10.03333 10.03333
#> p_best
#> D 0.90775
#> C 0.03975
#> B 0.00825
#> A 0.04425
nma_mv$fit_stats
#> model_type heterogeneity n_treatments n_edges estimator QE QE_df
#> 1 <NA> <NA> 9 6 MLE 9.140389 6
#> QE_p logLik Dev AIC BIC AICc
#> 1 0.1658353 7.234778 9.530444 13.53044 20.31914 -126.4696
nma_mv$level_heterogeneity_summary
#> NULL
nma_mv$heterogeneity_summary$level_heterogeneity
#> NULLMultivariate Variants
For multivariate NMA where studies contribute multiple outcomes or
multiple contrasts, cluster-robust SEs can be requested by setting
robustID:
nma_mv_cluster <- suppressWarnings(network_meta(
data = nma_dat,
study_id = "study",
treatment_1 = "trt1",
treatment_2 = "trt2",
effect = "yi",
variance = "vi",
reference = "A",
model_type = "multivariate",
heterogeneity = "flexible",
moderators = c("severity", "followup_months"),
estimation_method = "MLE",
within_varcov_type = "multilevel",
robustID = "study"
))
nma_mv_cluster$fit_coefficients
nma_mv_cluster$robust_cluster_id
network_forest_plot(nma_mv, effect_type = "total", main = "Multivariate: Total")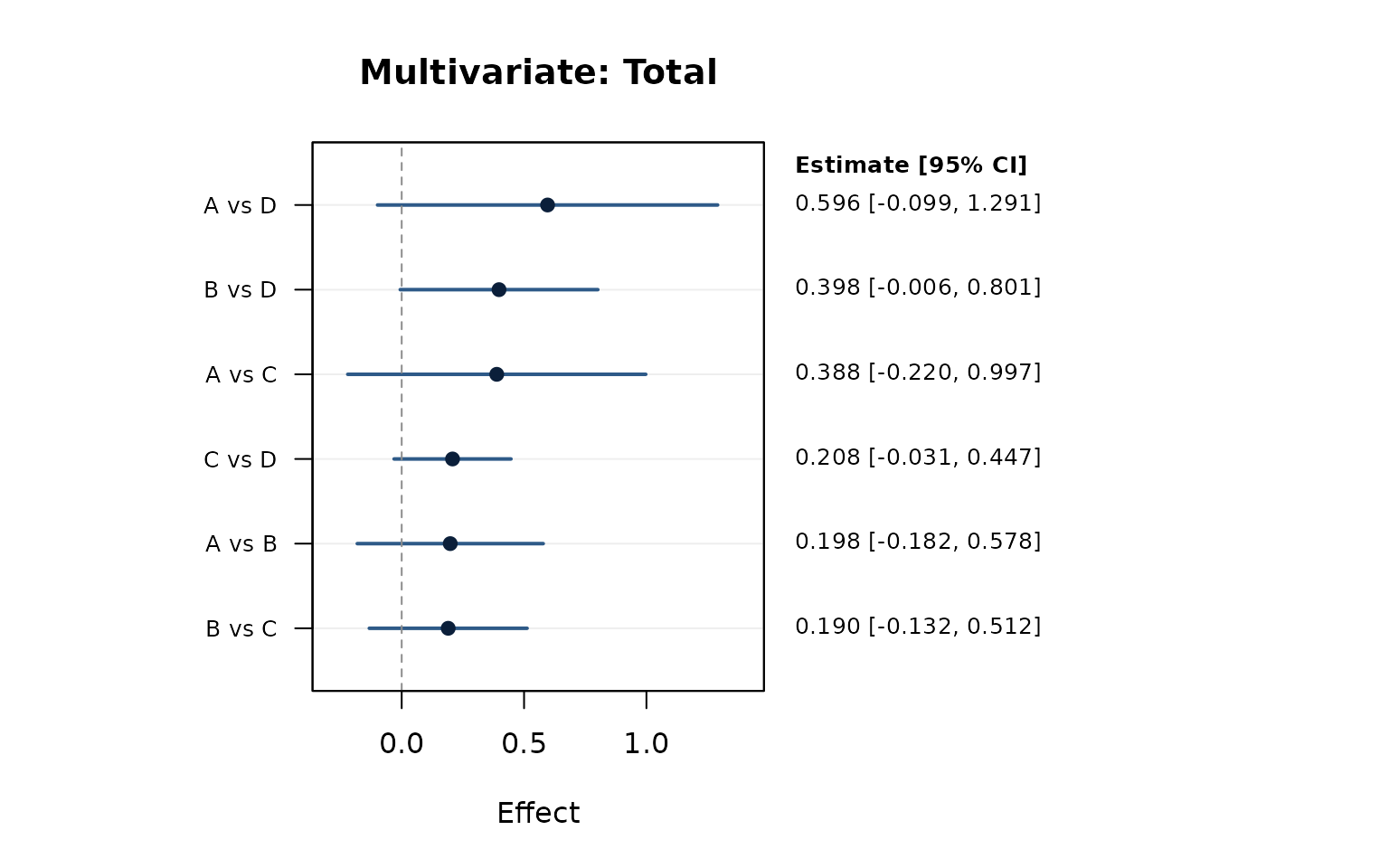
network_forest_overlay_plot(nma_mv, order_by = "total", right_label = "total",
main = "Multivariate: Overlay")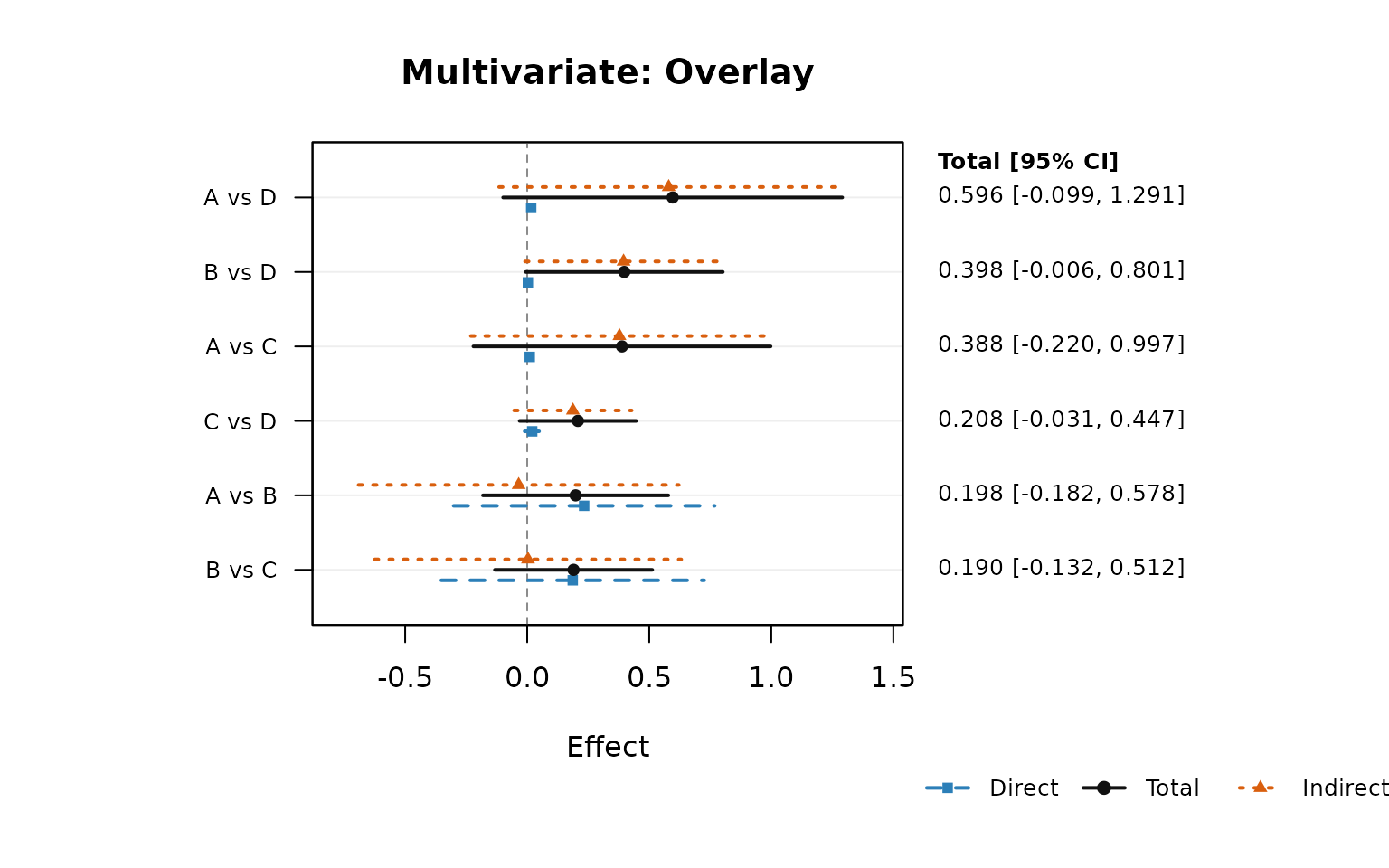
network_contribution_heatmap(nma_mv, main = "Multivariate: Contribution")
Compare multilevel and multivariate model fit side-by-side on the same dataset
mars:::compare_nma_models(
multilevel = nma_ml,
multivariate = nma_mv
)
#> model_type heterogeneity n_treatments n_edges estimator QE
#> multilevel multilevel flexible 4 6 MLE 0.106132
#> multivariate multivariate flexible 4 6 MLE 9.140389
#> QE_df QE_p logLik Dev AIC BIC AICc
#> multilevel 7 0.9999972 3.609517 16.780967 6.780967 10.17531 -133.2190
#> multivariate 6 0.1658353 7.234778 9.530444 13.530444 20.31914 -126.4696
#> AIC_rank BIC_rank AICc_rank
#> multilevel 1 1 1
#> multivariate 2 2 2