
Publication Bias Models with MARS
mars authors
2026-05-15
Publication-Bias-Models.RmdThis vignette demonstrates publication-bias modeling with
publication_bias(method = "beta_selection") and
publication_bias(method = "beta_binomial") and
publication_bias(method = "logistic_selection") after
fitting mars models.
Current implementation note: - Univariate publication-bias fits
estimate a scalar tau2 inside the selection model. -
Multilevel and multivariate publication-bias fits use the fitted MARS
covariance blocks directly, rather than collapsing to diagonal sampling
variances. - Reported standard errors use a conservative rule: when both
Hessian-based and OPG-based SEs are finite, the larger value is
reported.
library(mars)
extract_full_data_coefficients <- function(fit) {
est_raw <- if(!is.null(fit$beta_r)) {
fit$beta_r
} else if(!is.null(fit$beta_fe)) {
fit$beta_fe
} else {
fit$est_values$mu
}
est_vec <- as.numeric(est_raw)
est_names <- names(est_raw)
if(is.matrix(est_raw) || is.data.frame(est_raw)) {
est_vec <- as.numeric(est_raw[, 1, drop = TRUE])
est_names <- rownames(est_raw)
}
if(length(est_names) != length(est_vec)) {
est_names <- colnames(fit$design_matrix)
}
se_vec <- rep(NA_real_, length(est_vec))
vc <- fit$varcov_beta
if(is.matrix(vc) && nrow(vc) == length(est_vec)) {
se_vec <- sqrt(pmax(diag(vc), 0))
if(!is.null(colnames(vc)) && length(colnames(vc)) == length(est_vec)) {
est_names <- colnames(vc)
}
}
data.frame(
term = est_names,
estimate = est_vec,
std_error = se_vec,
stringsAsFactors = FALSE
)
}
compare_pub_bias_3 <- function(x, y, z,
label_x = "beta_selection",
label_y = "beta_binomial",
label_z = "logistic_selection",
ref = NULL,
label_ref = "all_data") {
a <- x$coefficients
b <- y$coefficients
c <- z$coefficients
names(a)[names(a) == "estimate"] <- paste0("estimate_", label_x)
names(a)[names(a) == "std_error"] <- paste0("se_", label_x)
names(b)[names(b) == "estimate"] <- paste0("estimate_", label_y)
names(b)[names(b) == "std_error"] <- paste0("se_", label_y)
names(c)[names(c) == "estimate"] <- paste0("estimate_", label_z)
names(c)[names(c) == "std_error"] <- paste0("se_", label_z)
out <- merge(
a[, c("term", paste0("estimate_", label_x), paste0("se_", label_x))],
b[, c("term", paste0("estimate_", label_y), paste0("se_", label_y))],
by = "term",
all = TRUE
)
out <- merge(
out,
c[, c("term", paste0("estimate_", label_z), paste0("se_", label_z))],
by = "term",
all = TRUE
)
if(!is.null(ref)) {
r <- ref
names(r)[names(r) == "estimate"] <- paste0("estimate_", label_ref)
names(r)[names(r) == "std_error"] <- paste0("se_", label_ref)
out <- merge(
out,
r[, c("term", paste0("estimate_", label_ref), paste0("se_", label_ref))],
by = "term",
all = TRUE
)
}
out[order(out$term), ]
}
plot_pub_bias_compare_3 <- function(tbl,
label_x = "beta_selection",
label_y = "beta_binomial",
label_z = "logistic_selection",
label_ref = "all_data",
main = "Method Comparison") {
ref_est_name <- paste0("estimate_", label_ref)
ref_se_name <- paste0("se_", label_ref)
est_x <- tbl[[paste0("estimate_", label_x)]]
se_x <- tbl[[paste0("se_", label_x)]]
est_y <- tbl[[paste0("estimate_", label_y)]]
se_y <- tbl[[paste0("se_", label_y)]]
est_z <- tbl[[paste0("estimate_", label_z)]]
se_z <- tbl[[paste0("se_", label_z)]]
has_ref <- ref_est_name %in% names(tbl)
est_ref <- if(has_ref) tbl[[ref_est_name]] else rep(NA_real_, nrow(tbl))
se_ref <- if(has_ref && ref_se_name %in% names(tbl)) tbl[[ref_se_name]] else rep(NA_real_, nrow(tbl))
k <- nrow(tbl)
xpos <- seq_len(k)
offsets <- if(has_ref) c(-0.27, -0.09, 0.09, 0.27) else c(-0.18, 0, 0.18)
all_vals <- c(
est_x, est_y, est_z, est_ref,
est_x - 1.96 * se_x, est_x + 1.96 * se_x,
est_y - 1.96 * se_y, est_y + 1.96 * se_y,
est_z - 1.96 * se_z, est_z + 1.96 * se_z,
est_ref - 1.96 * se_ref, est_ref + 1.96 * se_ref
)
ylim <- range(all_vals[is.finite(all_vals)])
old_par <- par(no.readonly = TRUE)
on.exit(par(old_par), add = TRUE)
x_axis_cex <- max(0.55, min(0.8, 8 / max(nchar(tbl$term), 1L)))
mar <- old_par$mar
mar[1] <- max(mar[1], 8 + 3 * x_axis_cex)
par(mar = mar)
plot(xpos, est_x, type = "n", xaxt = "n", xlab = "", ylab = "Estimate", main = main, ylim = ylim)
abline(h = 0, lty = 2, col = "gray60")
axis(1, at = xpos, labels = tbl$term, las = 2, cex.axis = x_axis_cex)
title(xlab = "Coefficient Term", line = 6)
keep_x <- is.finite(est_x) & is.finite(se_x)
keep_y <- is.finite(est_y) & is.finite(se_y)
keep_z <- is.finite(est_z) & is.finite(se_z)
keep_ref <- has_ref & is.finite(est_ref) & is.finite(se_ref)
pt_x <- is.finite(est_x)
pt_y <- is.finite(est_y)
pt_z <- is.finite(est_z)
pt_ref <- has_ref & is.finite(est_ref)
miss_se_x <- is.finite(est_x) & !is.finite(se_x)
miss_se_y <- is.finite(est_y) & !is.finite(se_y)
miss_se_z <- is.finite(est_z) & !is.finite(se_z)
miss_se_ref <- has_ref & is.finite(est_ref) & !is.finite(se_ref)
x_x <- xpos + offsets[1]
x_y <- xpos + offsets[2]
x_z <- xpos + offsets[3]
x_ref <- if(has_ref) xpos + offsets[4] else xpos
if(any(keep_x)) {
segments(x_x[keep_x], est_x[keep_x] - 1.96 * se_x[keep_x],
x_x[keep_x], est_x[keep_x] + 1.96 * se_x[keep_x], col = "#1F78B4", lwd = 2)
}
if(any(keep_y)) {
segments(x_y[keep_y], est_y[keep_y] - 1.96 * se_y[keep_y],
x_y[keep_y], est_y[keep_y] + 1.96 * se_y[keep_y], col = "#E31A1C", lwd = 2)
}
if(any(keep_z)) {
segments(x_z[keep_z], est_z[keep_z] - 1.96 * se_z[keep_z],
x_z[keep_z], est_z[keep_z] + 1.96 * se_z[keep_z], col = "#33A02C", lwd = 2)
}
if(any(keep_ref)) {
segments(x_ref[keep_ref], est_ref[keep_ref] - 1.96 * se_ref[keep_ref],
x_ref[keep_ref], est_ref[keep_ref] + 1.96 * se_ref[keep_ref], col = "#4D4D4D", lwd = 2)
}
if(any(pt_x)) {
points(x_x[pt_x], est_x[pt_x], pch = 19, col = "#1F78B4")
}
if(any(pt_y)) {
points(x_y[pt_y], est_y[pt_y], pch = 17, col = "#E31A1C")
}
if(any(pt_z)) {
points(x_z[pt_z], est_z[pt_z], pch = 15, col = "#33A02C")
}
if(any(pt_ref)) {
points(x_ref[pt_ref], est_ref[pt_ref], pch = 18, col = "#4D4D4D")
}
if(any(miss_se_x)) {
points(x_x[miss_se_x], est_x[miss_se_x], pch = 1, col = "#1F78B4", cex = 1.2, lwd = 1.5)
}
if(any(miss_se_y)) {
points(x_y[miss_se_y], est_y[miss_se_y], pch = 2, col = "#E31A1C", cex = 1.2, lwd = 1.5)
}
if(any(miss_se_z)) {
points(x_z[miss_se_z], est_z[miss_se_z], pch = 0, col = "#33A02C", cex = 1.2, lwd = 1.5)
}
if(any(miss_se_ref)) {
points(x_ref[miss_se_ref], est_ref[miss_se_ref], pch = 5, col = "#4D4D4D", cex = 1.2, lwd = 1.5)
}
legend_labels <- c(label_x, label_y, label_z)
legend_cols <- c("#1F78B4", "#E31A1C", "#33A02C")
legend_pch <- c(19, 17, 15)
if(has_ref) {
legend_labels <- c(legend_labels, label_ref)
legend_cols <- c(legend_cols, "#4D4D4D")
legend_pch <- c(legend_pch, 18)
}
legend("topright", legend = legend_labels, col = legend_cols, pch = legend_pch, bty = "n")
if(any(miss_se_x) || any(miss_se_y) || any(miss_se_z) || any(miss_se_ref)) {
mtext("Open symbols indicate estimates with unavailable SE.", side = 3, line = 0.2, cex = 0.8)
}
}
extract_alt_interval <- function(x, term = "(Intercept)") {
if(!is.null(x$summary) &&
all(c("point_estimate", "ci_lower", "ci_upper") %in% colnames(x$summary)) &&
term %in% rownames(x$summary)) {
out <- x$summary[term, c("point_estimate", "ci_lower", "ci_upper"), drop = FALSE]
return(out)
}
if(!is.null(x$summary) &&
all(c("estimate", "ci_lower", "ci_upper") %in% colnames(x$summary)) &&
term %in% rownames(x$summary)) {
out <- x$summary[term, c("estimate", "ci_lower", "ci_upper"), drop = FALSE]
names(out) <- c("point_estimate", "ci_lower", "ci_upper")
return(out)
}
stop("Could not extract a point estimate + interval for term: ", term)
}Univariate Example
fit_uni <- mars(
formula = yi ~ 1,
data = bcg,
studyID = "trial",
variance = "vi",
varcov_type = "univariate",
structure = "univariate",
estimation_method = "MLE"
)
summary(fit_uni)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> univariate
#>
#> Estimation Method:
#> Maximum Likelihood
#>
#> Model Formula:
#> yi ~ 1
#>
#> Data Summary:
#> Number of Effect Sizes: 13
#> Number of Fixed Effects: 1
#> Number of Random Effects: 1
#>
#> Random Components:
#> term var SD
#> intercept 0.28 0.5292
#>
#> Fixed Effects Estimates:
#> attribute estimate SE z_test p_value lower upper
#> (Intercept) -0.7112 0.1719 -4.137 3.513e-05 -1.048 -0.3743
#>
#> Model Fit Statistics:
#> logLik Dev AIC BIC AICc
#> -12.67 31.33 29.33 30.46 37.9
#>
#> Q Error: 152.233 (12), p < 0.0001
#>
#> I2 (General): 92.03957
#>
#> Residual Diagnostics:
#> n n_finite_raw mean_raw sd_raw rmse mae q_pearson mean_abs_studentized
#> 13 13 -0.02945 0.6938 0.6672 0.5958 13.16 0.9527
#> max_abs_studentized prop_abs_studentized_gt2 prop_abs_studentized_gt3
#> 1.434 0 0
#>
#> Normality (whitened residuals): test n_tested statistic p_value
#> shapiro_wilk_whitened 13 0.886 0.08592
#>
#> Heteroscedasticity trend (|raw residual| ~ fitted): n corr_abs_raw_fitted slope p_value
#> 13 NA NA NA
pb_uni <- publication_bias(
fit_uni,
method = "beta_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_uni)
#> Publication Bias Model (MARS)
#> Method: beta_selection
#> Structure: univariate
#> Effects: 13 | Studies: 13
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: 9.147
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 1.1797
#> beta 0.0008
#> tau2 0.4513
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> beta_selection 0.0100 0.4415
#> beta_selection 0.1189 0.7740
#> beta_selection 0.2278 0.9924
#> beta_selection 0.3367 1.2392
#> beta_selection 0.4456 1.5590
#> beta_selection 0.5544 2.0173
#> beta_selection 0.6633 2.7566
#> beta_selection 0.7722 4.1858
#> beta_selection 0.8811 8.2075
#> beta_selection 0.9900 99.4385
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) -1.3572 0.252 -5.3848 0
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) -0.7112 0.1719 -1.0481
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> -0.3743 -1.3572 0.252 -1.8512 -0.8632
#> estimate_diff percent_change direction_changed ci_overlap
#> -0.646 -90.8364 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -7.8519 SE: 3.2091 t: -2.4468 df: 12 p: 0.0308
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one.
pb_uni_bb <- publication_bias(
fit_uni,
method = "beta_binomial",
n_bins = 8,
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_uni_bb)
#> Publication Bias Model (MARS)
#> Method: beta_binomial
#> Structure: univariate
#> Effects: 13 | Studies: 13
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -33.37
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 7213.1265
#> beta 0.0006
#> tau2 1.6265
#> n_bins 8.0000
#>
#> Selection Weight Profile (bin probabilities):
#> bin p_lower p_upper weight
#> 1 0.000 0.125 0
#> 2 0.125 0.250 0
#> 3 0.250 0.375 0
#> 4 0.375 0.500 0
#> 5 0.500 0.625 0
#> 6 0.625 0.750 0
#> 7 0.750 0.875 0
#> 8 0.875 1.000 1
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) 0.3576 0.4306 0.8304 0.4063
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) -0.7112 0.1719 -1.0481
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> -0.3743 0.3576 0.4306 -0.4864 1.2016
#> estimate_diff percent_change direction_changed ci_overlap
#> 1.0688 150.2788 TRUE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -7.8519 SE: 3.2091 t: -2.4468 df: 12 p: 0.0308
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. beta_binomial uses discretized p-value bins; coefficient magnitudes can differ materially from beta_selection.
pb_uni_logit <- publication_bias(
fit_uni,
method = "logistic_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_uni_logit)
#> Publication Bias Model (MARS)
#> Method: logistic_selection
#> Structure: univariate
#> Effects: 13 | Studies: 13
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: 13.92
#>
#> Selection Parameters:
#> parameter estimate
#> logit_intercept 8.0045
#> logit_slope 0.5688
#> tau2 0.4256
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> logistic_selection 0.0100 1.0000
#> logistic_selection 0.1189 0.9999
#> logistic_selection 0.2278 0.9999
#> logistic_selection 0.3367 0.9998
#> logistic_selection 0.4456 0.9998
#> logistic_selection 0.5544 0.9998
#> logistic_selection 0.6633 0.9997
#> logistic_selection 0.7722 0.9997
#> logistic_selection 0.8811 0.9997
#> logistic_selection 0.9900 0.9997
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) -0.7723 0.2848 -2.7119 0.0067
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) -0.7112 0.1719 -1.0481
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> -0.3743 -0.7723 0.2848 -1.3305 -0.2141
#> estimate_diff percent_change direction_changed ci_overlap
#> -0.0611 -8.5908 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -7.8519 SE: 3.2091 t: -2.4468 df: 12 p: 0.0308
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one.Selection and adjusted fixed effects:
pb_uni$selection
#> alpha beta tau2
#> 1.1797078025 0.0008305913 0.4512552801
pb_uni$covariance_mode
#> [1] "publication_bias_tau2"
pb_uni$coefficients
#> term estimate std_error z_value p_value
#> 1 (Intercept) -1.357227 0.25205 -5.384754 7.254387e-08
pb_uni_bb$selection_weights
#> bin p_lower p_upper weight
#> 1 1 0.000 0.125 4.187106e-28
#> 2 2 0.125 0.250 3.523234e-24
#> 3 3 0.250 0.375 1.524843e-20
#> 4 4 0.375 0.500 4.583461e-17
#> 5 5 0.500 0.625 1.102277e-13
#> 6 6 0.625 0.750 2.385876e-10
#> 7 7 0.750 0.875 5.737124e-07
#> 8 8 0.875 1.000 9.999994e-01
pb_uni_logit$selection
#> logit_intercept logit_slope tau2
#> 8.0045371 0.5687618 0.4255908
pb_uni_logit$coefficients
#> term estimate std_error z_value p_value
#> 1 (Intercept) -0.7722971 0.2847817 -2.711892 0.006690042For the univariate case, the publication-bias likelihood still
estimates a scalar tau2, so the selection parameter vector
includes tau2 along with the selection-function
parameters.
ref_uni <- extract_full_data_coefficients(fit_uni)
cmp_uni <- compare_pub_bias_3(pb_uni, pb_uni_bb, pb_uni_logit, ref = ref_uni)
cmp_uni
#> term estimate_beta_selection se_beta_selection estimate_beta_binomial
#> 1 (Intercept) -1.357227 0.25205 0.3575826
#> se_beta_binomial estimate_logistic_selection se_logistic_selection
#> 1 0.4306209 -0.7722971 0.2847817
#> estimate_all_data se_all_data
#> 1 -0.7111991 0.1718968
plot_pub_bias_compare_3(cmp_uni, main = "Univariate: publication-bias methods with all-data reference")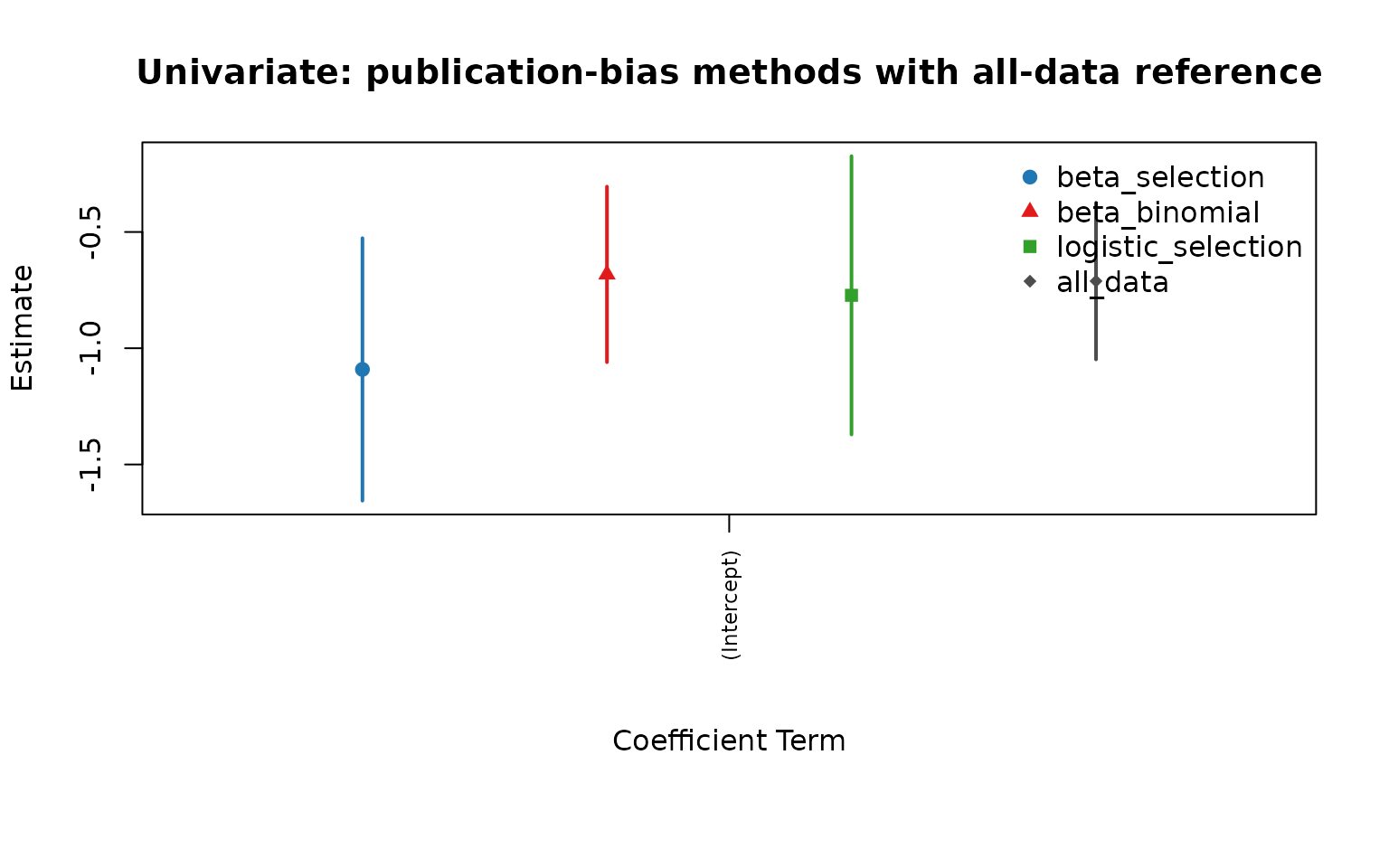
Funnel plot from the publication-bias fit:
funnel_plot(pb_uni)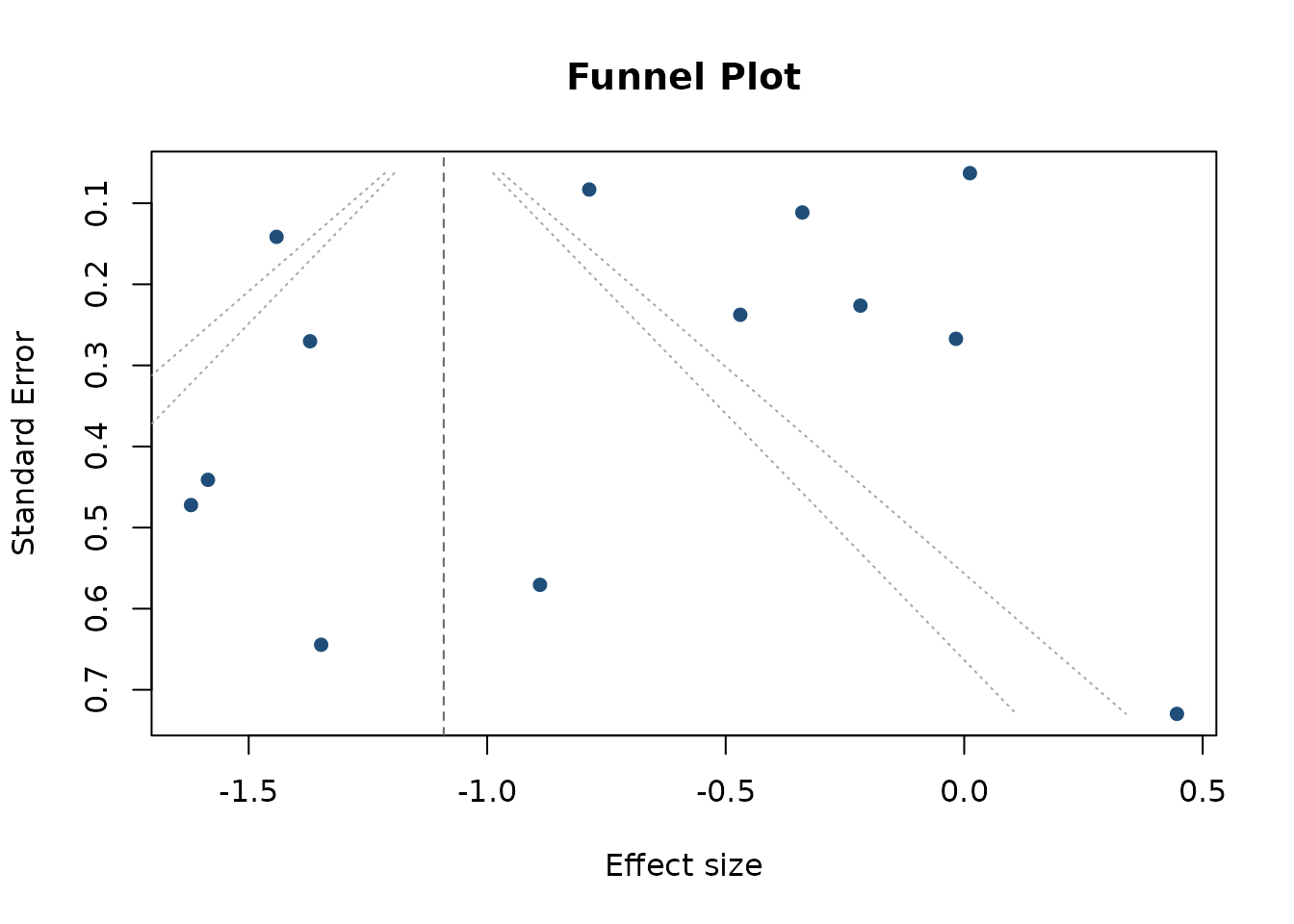
Jackknife version of the complete-data fit and each publication-bias model:
set.seed(2026)
# Keep the vignette jackknife examples intentionally small/lightweight.
bcg_jack <- bcg[seq_len(min(6, nrow(bcg))), , drop = FALSE]
jack_complete <- mars_alt_estimation(
type = "jackknife",
data = bcg_jack,
formula = yi ~ 1,
studyID = "trial",
effectID = NULL,
sample_size = NULL,
variance = "vi",
varcov_type = "univariate",
structure = "univariate",
estimation_method = "MLE",
jackknife_target = "complete_data"
)
jack_beta_selection <- mars_alt_estimation(
type = "jackknife",
data = bcg_jack,
formula = yi ~ 1,
studyID = "trial",
effectID = NULL,
sample_size = NULL,
variance = "vi",
varcov_type = "univariate",
structure = "univariate",
estimation_method = "MLE",
jackknife_target = "beta_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
)
jack_beta_binomial <- mars_alt_estimation(
type = "jackknife",
data = bcg_jack,
formula = yi ~ 1,
studyID = "trial",
effectID = NULL,
sample_size = NULL,
variance = "vi",
varcov_type = "univariate",
structure = "univariate",
estimation_method = "MLE",
jackknife_target = "beta_binomial",
publication_bias_args = list(
p_value_tail = "two.sided",
n_bins = 8,
maxit = 100,
integration_points = 31
)
)
jack_logistic <- mars_alt_estimation(
type = "jackknife",
data = bcg_jack,
formula = yi ~ 1,
studyID = "trial",
effectID = NULL,
sample_size = NULL,
variance = "vi",
varcov_type = "univariate",
structure = "univariate",
estimation_method = "MLE",
jackknife_target = "logistic_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
)
jack_complete_stats <- compute_alt_stats(jack_complete, type = "jackknife")
jack_beta_selection_stats <- compute_alt_stats(jack_beta_selection, type = "jackknife")
jack_beta_binomial_stats <- compute_alt_stats(jack_beta_binomial, type = "jackknife")
jack_logistic_stats <- compute_alt_stats(jack_logistic, type = "jackknife")
jackknife_compare <- rbind(
complete_data = extract_alt_interval(jack_complete_stats),
beta_selection = extract_alt_interval(jack_beta_selection_stats),
beta_binomial = extract_alt_interval(jack_beta_binomial_stats),
logistic_selection = extract_alt_interval(jack_logistic_stats)
)
jackknife_compareThe univariate jackknife illustration uses only the first six trials so the vignette demonstrates the workflow without refitting all leave-one-out models from the full dataset.
jack_est <- jackknife_compare[, "point_estimate"]
jack_lo <- jackknife_compare[, "ci_lower"]
jack_hi <- jackknife_compare[, "ci_upper"]
xpos <- seq_len(nrow(jackknife_compare))
plot(
xpos, jack_est,
ylim = range(c(jack_lo, jack_hi)),
xaxt = "n", xlab = "", ylab = "Jackknife Estimate",
pch = 19, col = "#1F78B4",
main = "Univariate Jackknife: complete data and publication-bias targets"
)
segments(xpos, jack_lo, xpos, jack_hi, lwd = 2, col = "#1F78B4")
abline(h = 0, lty = 2, col = "gray60")
axis(1, at = xpos, labels = rownames(jackknife_compare), las = 2)
title(xlab = "Jackknife Target", line = 6)Multilevel Example
fit_ml <- mars(
formula = effect ~ 1 + (1 | district/study),
data = school,
studyID = "district",
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE"
)
summary(fit_ml)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multilevel
#>
#> Estimation Method:
#> Maximum Likelihood
#>
#> Model Formula:
#> effect ~ 1 + (1 | district/study)
#>
#> Data Summary:
#> Number of Effect Sizes: 56
#> Number of Fixed Effects: 1
#> Number of Random Effects: 2
#>
#> Random Data Summary:
#> Number unique district: 11
#> Number unique study: 56
#>
#> Random Components:
#> term var SD
#> district 0.05774 0.2403
#> study 0.03286 0.1813
#>
#> Fixed Effects Estimates:
#> attribute estimate SE z_test p_value lower upper
#> (Intercept) 0.1845 0.08048 2.292 0.02191 0.02671 0.3422
#>
#> Model Fit Statistics:
#> logLik Dev AIC BIC AICc
#> -8.395 24.79 22.79 28.87 24.5
#>
#> Q Error: 578.864 (55), p < 0.0001
#>
#> I2 (General):
#> names values
#> district 92.38
#> study 87.34
#>
#> I2 (Jackson): 98.69621
#> I2 (Between): 86.3727
#>
#> Residual Diagnostics:
#> n n_finite_raw mean_raw sd_raw rmse mae q_pearson mean_abs_studentized
#> 56 56 -0.06446 0.3258 0.3293 0.2632 96.8 1.065
#> max_abs_studentized prop_abs_studentized_gt2 prop_abs_studentized_gt3
#> 4.196 0.1071 0.03571
#>
#> Normality (whitened residuals): test n_tested statistic p_value
#> shapiro_wilk_whitened 56 0.9555 0.03766
#>
#> Heteroscedasticity trend (|raw residual| ~ fitted): n corr_abs_raw_fitted slope p_value
#> 56 NA NA NA
pb_ml <- publication_bias(
fit_ml,
method = "beta_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_ml)
#> Publication Bias Model (MARS)
#> Method: beta_selection
#> Structure: multilevel
#> Effects: 56 | Studies: 11
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -142.5
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 2179.5386
#> beta 0.0003
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> beta_selection 0.0100 0
#> beta_selection 0.1189 0
#> beta_selection 0.2278 0
#> beta_selection 0.3367 0
#> beta_selection 0.4456 0
#> beta_selection 0.5544 0
#> beta_selection 0.6633 0
#> beta_selection 0.7722 0
#> beta_selection 0.8811 0
#> beta_selection 0.9900 0
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) 0.9456 0.0332 28.4619 0
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) 0.1845 0.0805 0.0267
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> 0.3422 0.9456 0.0332 0.8805 1.0107
#> estimate_diff percent_change direction_changed ci_overlap
#> 0.7611 412.6349 FALSE FALSE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: 3.69 SE: 1.8503 t: 1.9943 df: 10 p: 0.0741
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood.
pb_ml_bb <- publication_bias(
fit_ml,
method = "beta_binomial",
n_bins = 8,
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_ml_bb)
#> Publication Bias Model (MARS)
#> Method: beta_binomial
#> Structure: multilevel
#> Effects: 56 | Studies: 11
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -400.3
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 158.6092
#> beta 0.8091
#> n_bins 8.0000
#>
#> Selection Weight Profile (bin probabilities):
#> bin p_lower p_upper weight
#> 1 0.000 0.125 0.0000
#> 2 0.125 0.250 0.0000
#> 3 0.250 0.375 0.0000
#> 4 0.375 0.500 0.0000
#> 5 0.500 0.625 0.0000
#> 6 0.625 0.750 0.0011
#> 7 0.750 0.875 0.0332
#> 8 0.875 1.000 0.9656
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) 1.2761 0.034 37.5353 0
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) 0.1845 0.0805 0.0267
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> 0.3422 1.2761 0.034 1.2094 1.3427
#> estimate_diff percent_change direction_changed ci_overlap
#> 1.0916 591.7976 FALSE FALSE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: 3.69 SE: 1.8503 t: 1.9943 df: 10 p: 0.0741
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood. beta_binomial uses discretized p-value bins; coefficient magnitudes can differ materially from beta_selection.
pb_ml_logit <- publication_bias(
fit_ml,
method = "logistic_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_ml_logit)
#> Publication Bias Model (MARS)
#> Method: logistic_selection
#> Structure: multilevel
#> Effects: 56 | Studies: 11
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: 8.396
#>
#> Selection Parameters:
#> parameter estimate
#> logit_intercept 7.8501
#> logit_slope 11.7805
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> logistic_selection 0.0100 1.0000
#> logistic_selection 0.1189 1.0000
#> logistic_selection 0.2278 1.0000
#> logistic_selection 0.3367 1.0000
#> logistic_selection 0.4456 1.0000
#> logistic_selection 0.5544 1.0000
#> logistic_selection 0.6633 1.0000
#> logistic_selection 0.7722 1.0000
#> logistic_selection 0.8811 0.9999
#> logistic_selection 0.9900 0.9997
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> (Intercept) 0.1844 0.087 2.1183 0.0341
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se unadjusted_ci_lower
#> (Intercept) 0.1845 0.0805 0.0267
#> unadjusted_ci_upper estimate std_error adjusted_ci_lower adjusted_ci_upper
#> 0.3422 0.1844 0.087 0.0138 0.355
#> estimate_diff percent_change direction_changed ci_overlap
#> -1e-04 -0.0299 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: 3.69 SE: 1.8503 t: 1.9943 df: 10 p: 0.0741
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood.Multilevel coefficients with standard errors:
pb_ml$covariance_mode
#> [1] "fixed_mars_structure"
pb_ml$covariance_structure
#> $structure
#> [1] "multilevel"
#>
#> $Tau
#> [,1] [,2]
#> [1,] 0.05773805 0.00000000
#> [2,] 0.00000000 0.03286473
pb_ml$coefficients
#> term estimate std_error z_value p_value
#> 1 (Intercept) 0.9455827 0.03322272 28.46192 3.469368e-178
pb_ml_bb$coefficients
#> term estimate std_error z_value p_value
#> 1 (Intercept) 1.276058 0.0339962 37.53531 2.446214e-308
pb_ml_logit$coefficients
#> term estimate std_error z_value p_value
#> 1 (Intercept) 0.1844002 0.08704963 2.118334 0.03414679For multilevel fits, publication_bias() now uses the
fitted MARS block covariance structure directly. In that case the output
stores covariance_mode = "fixed_mars_structure" and the
publication-bias selection parameters no longer include a separate
scalar tau2.
ref_ml <- extract_full_data_coefficients(fit_ml)
cmp_ml <- compare_pub_bias_3(pb_ml, pb_ml_bb, pb_ml_logit, ref = ref_ml)
cmp_ml
#> term estimate_beta_selection se_beta_selection estimate_beta_binomial
#> 1 (Intercept) 0.9455827 0.03322272 1.276058
#> se_beta_binomial estimate_logistic_selection se_logistic_selection
#> 1 0.0339962 0.1844002 0.08704963
#> estimate_all_data se_all_data
#> 1 0.1844554 0.0804815
plot_pub_bias_compare_3(cmp_ml, main = "Multilevel: publication-bias methods with all-data reference")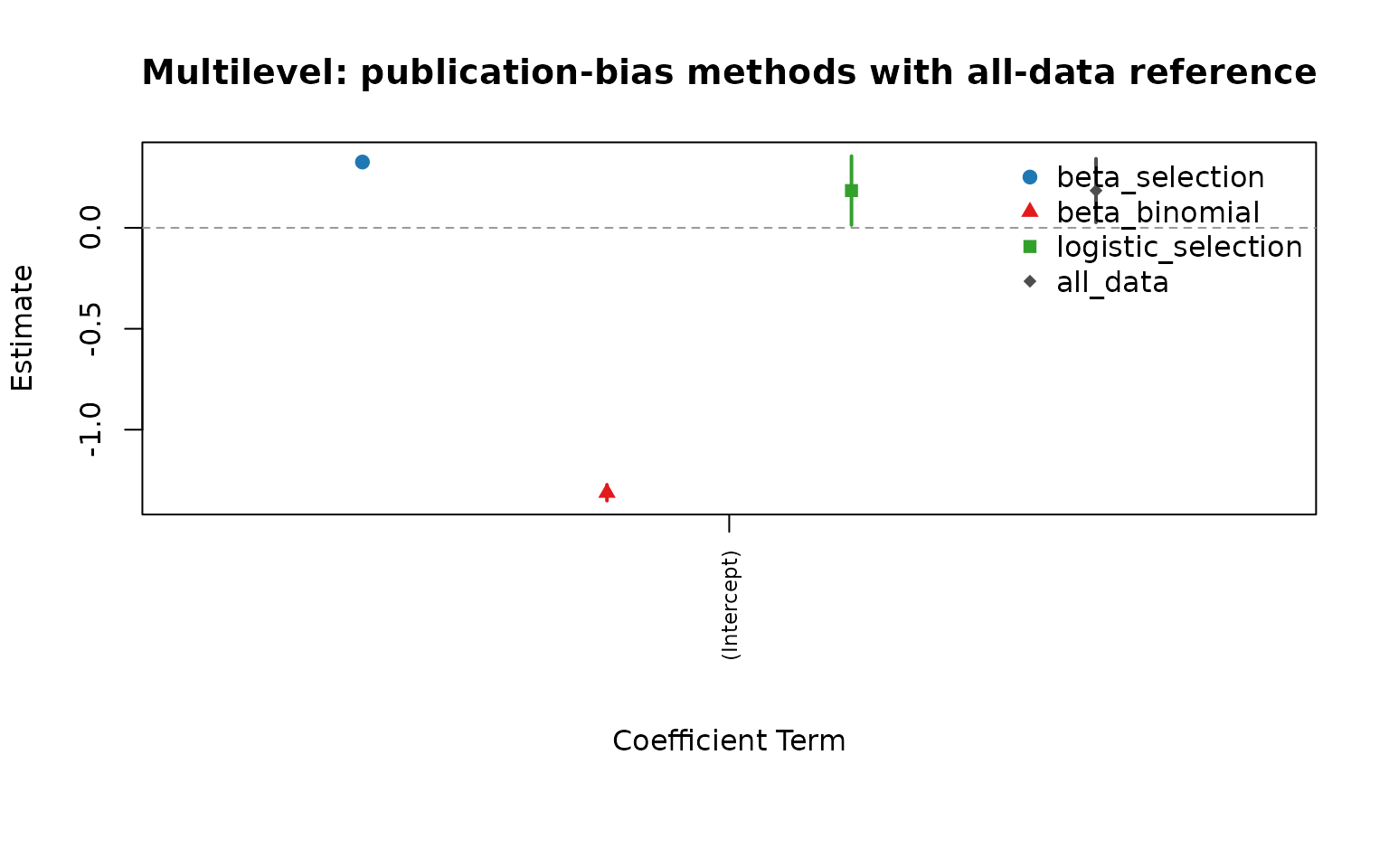
Jackknife version of the multilevel complete-data fit and each publication-bias model:
set.seed(2026)
school_jack <- subset(school, district %in% sort(unique(district))[1:4])
jack_ml_complete <- mars_alt_estimation(
type = "jackknife",
data = school_jack,
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE",
jackknife_target = "complete_data"
)
jack_ml_beta_selection <- mars_alt_estimation(
type = "jackknife",
data = school_jack,
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE",
jackknife_target = "beta_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 80,
integration_points = 31
)
)
jack_ml_beta_binomial <- mars_alt_estimation(
type = "jackknife",
data = school_jack,
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE",
jackknife_target = "beta_binomial",
publication_bias_args = list(
p_value_tail = "two.sided",
n_bins = 8,
maxit = 80,
integration_points = 31
)
)
jack_ml_logistic <- mars_alt_estimation(
type = "jackknife",
data = school_jack,
formula = effect ~ 1 + (1 | district/study),
studyID = "district",
effectID = NULL,
sample_size = NULL,
variance = "var",
varcov_type = "multilevel",
structure = "multilevel",
estimation_method = "MLE",
jackknife_target = "logistic_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 80,
integration_points = 31
)
)
jack_ml_complete_stats <- compute_alt_stats(jack_ml_complete, type = "jackknife")
jack_ml_beta_selection_stats <- compute_alt_stats(jack_ml_beta_selection, type = "jackknife")
jack_ml_beta_binomial_stats <- compute_alt_stats(jack_ml_beta_binomial, type = "jackknife")
jack_ml_logistic_stats <- compute_alt_stats(jack_ml_logistic, type = "jackknife")
jackknife_compare_ml <- rbind(
complete_data = extract_alt_interval(jack_ml_complete_stats),
beta_selection = extract_alt_interval(jack_ml_beta_selection_stats),
beta_binomial = extract_alt_interval(jack_ml_beta_binomial_stats),
logistic_selection = extract_alt_interval(jack_ml_logistic_stats)
)
jackknife_compare_mlThe multilevel jackknife illustration uses only the first four top-level clusters so the vignette stays quick to run while still showing the leave-one- cluster-out workflow.
jack_ml_est <- jackknife_compare_ml[, "point_estimate"]
jack_ml_lo <- jackknife_compare_ml[, "ci_lower"]
jack_ml_hi <- jackknife_compare_ml[, "ci_upper"]
xpos_ml <- seq_len(nrow(jackknife_compare_ml))
plot(
xpos_ml, jack_ml_est,
ylim = range(c(jack_ml_lo, jack_ml_hi)),
xaxt = "n", xlab = "", ylab = "Jackknife Estimate",
pch = 19, col = "#1F78B4",
main = "Multilevel Jackknife: complete data and publication-bias targets"
)
segments(xpos_ml, jack_ml_lo, xpos_ml, jack_ml_hi, lwd = 2, col = "#1F78B4")
abline(h = 0, lty = 2, col = "gray60")
axis(1, at = xpos_ml, labels = rownames(jackknife_compare_ml), las = 2)
title(xlab = "Jackknife Target", line = 6)Multivariate Example
becker09 <- na.omit(becker09)
fit_mv <- mars(
data = becker09,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
)
)
summary(fit_mv)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multivariate
#>
#> Estimation Method:
#> Restricted Maximum Likelihood
#>
#> Model Formula:
#> NULL
#>
#> Data Summary:
#> Number of Effect Sizes: 48
#> Number of Fixed Effects: 6
#> Number of Random Effects: 6
#>
#> Random Components:
#> ri_1 ri_2 ri_3 ri_4 ri_5 ri_6
#> ri_1 0.13204 0.07630 -0.03144 0.003058 -0.018150 0.0022236
#> ri_2 0.99878 0.04420 -0.01698 0.001532 -0.009861 0.0008355
#> ri_3 -0.42074 -0.39284 0.04229 -0.007313 0.009630 -0.0122546
#> ri_4 0.23176 0.20062 -0.97927 0.001319 -0.001498 0.0022776
#> ri_5 -0.62514 -0.58701 0.58603 -0.516444 0.006384 -0.0024641
#> ri_6 0.09632 0.06256 -0.93797 0.987249 -0.485426 0.0040360
#>
#> Fixed Effects Estimates:
#> attribute estimate SE z_test p_value lower upper
#> 1 -0.09773 0.13777 -0.7093 4.781e-01 -0.3677 0.172299
#> 2 -0.17556 0.08620 -2.0367 4.168e-02 -0.3445 -0.006613
#> 3 0.31868 0.08406 3.7911 1.500e-04 0.1539 0.483438
#> 4 0.52720 0.03335 15.8070 2.785e-56 0.4618 0.592564
#> 5 -0.41756 0.04590 -9.0968 9.305e-20 -0.5075 -0.327591
#> 6 -0.40071 0.04182 -9.5821 9.510e-22 -0.4827 -0.318750
#>
#> Model Fit Statistics:
#> logLik Dev AIC BIC AICc
#> 18.63 -37.25 16.75 63.67 29.48
#>
#> Q Error: 230.642 (42), p < 0.0001
#>
#> I2 (General):
#> names values
#> ri_1 94.53
#> ri_2 85.26
#> ri_3 84.69
#> ri_4 14.71
#> ri_5 45.51
#> ri_6 34.56
#>
#> I2 (Jackson):
#> names values
#> ri_1 90.98
#> ri_2 77.93
#> ri_3 81.33
#> ri_4 16.14
#> ri_5 43.25
#> ri_6 31.42
#>
#> I2 (Between): 83.393
#>
#> Residual Diagnostics:
#> n n_finite_raw mean_raw sd_raw rmse mae q_pearson mean_abs_studentized
#> 48 48 -0.02155 0.2127 0.2116 0.1646 133.5 1.419
#> max_abs_studentized prop_abs_studentized_gt2 prop_abs_studentized_gt3
#> 5.006 0.2708 0.1042
#>
#> Normality (whitened residuals): test n_tested statistic p_value
#> shapiro_wilk_whitened 48 0.9789 0.5351
#>
#> Heteroscedasticity trend (|raw residual| ~ fitted): n corr_abs_raw_fitted slope p_value
#> 48 -0.002893 -0.001091 0.9844
pb_mv <- publication_bias(
fit_mv,
method = "beta_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_mv)
#> Publication Bias Model (MARS)
#> Method: beta_selection
#> Structure: UN
#> Effects: 48 | Studies: 8
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -16.83
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 1.0114
#> beta 0.9479
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> beta_selection 0.0100 0.9493
#> beta_selection 0.1189 0.9825
#> beta_selection 0.2278 0.9966
#> beta_selection 0.3367 1.0090
#> beta_selection 0.4456 1.0217
#> beta_selection 0.5544 1.0360
#> beta_selection 0.6633 1.0534
#> beta_selection 0.7722 1.0769
#> beta_selection 0.8811 1.1157
#> beta_selection 0.9900 1.2710
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> factor(data[[effectID]])1 -0.1235 14.6343 -0.0084 0.9933
#> factor(data[[effectID]])2 -0.2511 10.8979 -0.0230 0.9816
#> factor(data[[effectID]])3 0.3092 3.3025 0.0936 0.9254
#> factor(data[[effectID]])4 0.5153 6.0255 0.0855 0.9318
#> factor(data[[effectID]])5 -0.4174 5.4063 -0.0772 0.9385
#> factor(data[[effectID]])6 -0.4235 2.8873 -0.1467 0.8834
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se
#> factor(data[[effectID]])1 -0.0977 0.1378
#> factor(data[[effectID]])2 -0.1756 0.0862
#> factor(data[[effectID]])3 0.3187 0.0841
#> factor(data[[effectID]])4 0.5272 0.0334
#> factor(data[[effectID]])5 -0.4176 0.0459
#> factor(data[[effectID]])6 -0.4007 0.0418
#> unadjusted_ci_lower unadjusted_ci_upper estimate std_error adjusted_ci_lower
#> -0.3678 0.1723 -0.1235 14.6343 -28.8066
#> -0.3445 -0.0066 -0.2511 10.8979 -21.6111
#> 0.1539 0.4834 0.3092 3.3025 -6.1636
#> 0.4618 0.5926 0.5153 6.0255 -11.2947
#> -0.5075 -0.3276 -0.4174 5.4063 -11.0137
#> -0.4827 -0.3187 -0.4235 2.8873 -6.0826
#> adjusted_ci_upper estimate_diff percent_change direction_changed ci_overlap
#> 28.5596 -0.0258 -26.3896 FALSE TRUE
#> 21.1088 -0.0756 -43.0508 FALSE TRUE
#> 6.7821 -0.0094 -2.9631 FALSE TRUE
#> 12.3253 -0.0119 -2.2526 FALSE TRUE
#> 10.1789 0.0001 0.0357 FALSE TRUE
#> 5.2355 -0.0228 -5.6967 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -3.8099 SE: 1.2322 t: -3.092 df: 7 p: 0.0175
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood.
pb_mv_bb <- publication_bias(
fit_mv,
method = "beta_binomial",
n_bins = 8,
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_mv_bb)
#> Publication Bias Model (MARS)
#> Method: beta_binomial
#> Structure: UN
#> Effects: 48 | Studies: 8
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -17.59
#>
#> Selection Parameters:
#> parameter estimate
#> alpha 1.5143
#> beta 1.4745
#> n_bins 8.0000
#>
#> Selection Weight Profile (bin probabilities):
#> bin p_lower p_upper weight
#> 1 0.000 0.125 0.0841
#> 2 0.125 0.250 0.1192
#> 3 0.250 0.375 0.1389
#> 4 0.375 0.500 0.1486
#> 5 0.500 0.625 0.1499
#> 6 0.625 0.750 0.1428
#> 7 0.750 0.875 0.1253
#> 8 0.875 1.000 0.0912
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> factor(data[[effectID]])1 -0.1111 0.2607 -0.4263 0.6699
#> factor(data[[effectID]])2 -0.2491 0.0691 -3.6043 0.0003
#> factor(data[[effectID]])3 0.3059 0.0907 3.3742 0.0007
#> factor(data[[effectID]])4 0.5172 0.4988 1.0369 0.2998
#> factor(data[[effectID]])5 -0.4182 0.2237 -1.8696 0.0615
#> factor(data[[effectID]])6 -0.4240 0.2348 -1.8058 0.0710
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se
#> factor(data[[effectID]])1 -0.0977 0.1378
#> factor(data[[effectID]])2 -0.1756 0.0862
#> factor(data[[effectID]])3 0.3187 0.0841
#> factor(data[[effectID]])4 0.5272 0.0334
#> factor(data[[effectID]])5 -0.4176 0.0459
#> factor(data[[effectID]])6 -0.4007 0.0418
#> unadjusted_ci_lower unadjusted_ci_upper estimate std_error adjusted_ci_lower
#> -0.3678 0.1723 -0.1111 0.2607 -0.6222
#> -0.3445 -0.0066 -0.2491 0.0691 -0.3845
#> 0.1539 0.4834 0.3059 0.0907 0.1282
#> 0.4618 0.5926 0.5172 0.4988 -0.4605
#> -0.5075 -0.3276 -0.4182 0.2237 -0.8567
#> -0.4827 -0.3187 -0.4240 0.2348 -0.8841
#> adjusted_ci_upper estimate_diff percent_change direction_changed ci_overlap
#> 0.3999 -0.0134 -13.7323 FALSE TRUE
#> -0.1136 -0.0735 -41.8828 FALSE TRUE
#> 0.4836 -0.0128 -4.0076 FALSE TRUE
#> 1.4949 -0.0100 -1.8977 FALSE TRUE
#> 0.0202 -0.0007 -0.1594 FALSE TRUE
#> 0.0362 -0.0232 -5.8017 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -3.8099 SE: 1.2322 t: -3.092 df: 7 p: 0.0175
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood. beta_binomial uses discretized p-value bins; coefficient magnitudes can differ materially from beta_selection.
pb_mv_logit <- publication_bias(
fit_mv,
method = "logistic_selection",
p_value_tail = "two.sided",
maxit = 100,
integration_points = 31
)
summary(pb_mv_logit)
#> Publication Bias Model (MARS)
#> Method: logistic_selection
#> Structure: UN
#> Effects: 48 | Studies: 8
#> P-value tail: two.sided
#> Convergence: 0 - Converged
#> Objective: -16.75
#>
#> Selection Parameters:
#> parameter estimate
#> logit_intercept 2.8831
#> logit_slope 21.4511
#>
#> Selection Weight Profile (p-value grid):
#> model p_value weight
#> logistic_selection 0.0100 1.0000
#> logistic_selection 0.1189 1.0000
#> logistic_selection 0.2278 1.0000
#> logistic_selection 0.3367 1.0000
#> logistic_selection 0.4456 1.0000
#> logistic_selection 0.5544 1.0000
#> logistic_selection 0.6633 1.0000
#> logistic_selection 0.7722 0.9998
#> logistic_selection 0.8811 0.9963
#> logistic_selection 0.9900 0.9568
#>
#> Bias-Adjusted Coefficients:
#> term estimate std_error z_value p_value
#> factor(data[[effectID]])1 -0.0974 0.2440 -0.3992 0.6897
#> factor(data[[effectID]])2 -0.2345 0.1673 -1.4017 0.1610
#> factor(data[[effectID]])3 0.2971 0.1265 2.3487 0.0188
#> factor(data[[effectID]])4 0.5148 0.0671 7.6661 0.0000
#> factor(data[[effectID]])5 -0.4170 0.0843 -4.9466 0.0000
#> factor(data[[effectID]])6 -0.4183 0.0630 -6.6361 0.0000
#>
#> Adjusted vs Unadjusted Summary:
#> term unadjusted_estimate unadjusted_se
#> factor(data[[effectID]])1 -0.0977 0.1378
#> factor(data[[effectID]])2 -0.1756 0.0862
#> factor(data[[effectID]])3 0.3187 0.0841
#> factor(data[[effectID]])4 0.5272 0.0334
#> factor(data[[effectID]])5 -0.4176 0.0459
#> factor(data[[effectID]])6 -0.4007 0.0418
#> unadjusted_ci_lower unadjusted_ci_upper estimate std_error adjusted_ci_lower
#> -0.3678 0.1723 -0.0974 0.2440 -0.5756
#> -0.3445 -0.0066 -0.2345 0.1673 -0.5625
#> 0.1539 0.4834 0.2971 0.1265 0.0492
#> 0.4618 0.5926 0.5148 0.0671 0.3832
#> -0.5075 -0.3276 -0.4170 0.0843 -0.5823
#> -0.4827 -0.3187 -0.4183 0.0630 -0.5418
#> adjusted_ci_upper estimate_diff percent_change direction_changed ci_overlap
#> 0.3808 0.0003 0.3268 FALSE TRUE
#> 0.0934 -0.0590 -33.6036 FALSE TRUE
#> 0.5451 -0.0215 -6.7566 FALSE TRUE
#> 0.6464 -0.0124 -2.3555 FALSE TRUE
#> -0.2518 0.0005 0.1270 FALSE TRUE
#> -0.2947 -0.0176 -4.3799 FALSE TRUE
#>
#> Small-Study Effects Check (Egger-type):
#> Model: SND ~ precision
#> Method: Egger-type small-study-effects regression with study-cluster robust SE (CR1)
#> Intercept: -3.8099 SE: 1.2322 t: -3.092 df: 7 p: 0.0175
#>
#> SE note:
#> SE rule: reported SEs use the larger of the Hessian-based and OPG-based values when both are finite, otherwise the finite one. Covariance structure: used the fitted MARS within/between-study covariance blocks directly for the publication-bias likelihood.Multivariate coefficients with standard errors:
pb_mv$covariance_mode
#> [1] "fixed_mars_structure"
pb_mv$covariance_structure
#> $structure
#> [1] "UN"
#>
#> $Tau
#> [,1] [,2] [,3] [,4] [,5]
#> [1,] 0.132040025 0.0763014664 -0.031441429 0.003058166 -0.018150222
#> [2,] 0.076301466 0.0441998006 -0.016984791 0.001531606 -0.009860779
#> [3,] -0.031441429 -0.0169847911 0.042292885 -0.007313107 0.009629524
#> [4,] 0.003058166 0.0015316061 -0.007313107 0.001318666 -0.001498453
#> [5,] -0.018150222 -0.0098607790 0.009629524 -0.001498453 0.006384168
#> [6,] 0.002223569 0.0008355477 -0.012254561 0.002277566 -0.002464062
#> [,6]
#> [1,] 0.0022235691
#> [2,] 0.0008355477
#> [3,] -0.0122545613
#> [4,] 0.0022775661
#> [5,] -0.0024640625
#> [6,] 0.0040360241
pb_mv$coefficients
#> term estimate std_error z_value p_value
#> 1 factor(data[[effectID]])1 -0.1235143 14.634251 -0.008440084 0.9932659
#> 2 factor(data[[effectID]])2 -0.2511335 10.897949 -0.023044110 0.9816151
#> 3 factor(data[[effectID]])3 0.3092381 3.302479 0.093638194 0.9253966
#> 4 factor(data[[effectID]])4 0.5153197 6.025506 0.085523056 0.9318456
#> 5 factor(data[[effectID]])5 -0.4174073 5.406261 -0.077208142 0.9384580
#> 6 factor(data[[effectID]])6 -0.4235416 2.887272 -0.146692669 0.8833746
pb_mv_bb$coefficients
#> term estimate std_error z_value p_value
#> 1 factor(data[[effectID]])1 -0.1111449 0.26074401 -0.4262606 0.6699179282
#> 2 factor(data[[effectID]])2 -0.2490830 0.06910738 -3.6042898 0.0003130077
#> 3 factor(data[[effectID]])3 0.3059095 0.09066155 3.3741922 0.0007403264
#> 4 factor(data[[effectID]])4 0.5171911 0.49880619 1.0368579 0.2998020984
#> 5 factor(data[[effectID]])5 -0.4182222 0.22369983 -1.8695689 0.0615437017
#> 6 factor(data[[effectID]])6 -0.4239621 0.23478051 -1.8057805 0.0709526289
pb_mv_logit$coefficients
#> term estimate std_error z_value p_value
#> 1 factor(data[[effectID]])1 -0.0974057 0.24397507 -0.3992445 6.897131e-01
#> 2 factor(data[[effectID]])2 -0.2345485 0.16733216 -1.4016944 1.610065e-01
#> 3 factor(data[[effectID]])3 0.2971491 0.12651639 2.3487004 1.883906e-02
#> 4 factor(data[[effectID]])4 0.5147775 0.06714979 7.6661065 1.772960e-14
#> 5 factor(data[[effectID]])5 -0.4170261 0.08430487 -4.9466427 7.550438e-07
#> 6 factor(data[[effectID]])6 -0.4182650 0.06302876 -6.6360968 3.220983e-11The same idea applies to multivariate fits: the publication-bias
likelihood is evaluated using the fitted MARS within-study covariance
blocks and the fitted between-study Tau matrix, rather than
replacing them with a single scalar heterogeneity term.
ref_mv <- extract_full_data_coefficients(fit_mv)
cmp_mv <- compare_pub_bias_3(pb_mv, pb_mv_bb, pb_mv_logit, ref = ref_mv)
cmp_mv
#> term estimate_beta_selection se_beta_selection
#> 1 factor(data[[effectID]])1 -0.1235143 14.634251
#> 2 factor(data[[effectID]])2 -0.2511335 10.897949
#> 3 factor(data[[effectID]])3 0.3092381 3.302479
#> 4 factor(data[[effectID]])4 0.5153197 6.025506
#> 5 factor(data[[effectID]])5 -0.4174073 5.406261
#> 6 factor(data[[effectID]])6 -0.4235416 2.887272
#> estimate_beta_binomial se_beta_binomial estimate_logistic_selection
#> 1 -0.1111449 0.26074401 -0.0974057
#> 2 -0.2490830 0.06910738 -0.2345485
#> 3 0.3059095 0.09066155 0.2971491
#> 4 0.5171911 0.49880619 0.5147775
#> 5 -0.4182222 0.22369983 -0.4170261
#> 6 -0.4239621 0.23478051 -0.4182650
#> se_logistic_selection estimate_all_data se_all_data
#> 1 0.24397507 -0.09772505 0.13776971
#> 2 0.16733216 -0.17555552 0.08619682
#> 3 0.12651639 0.31868112 0.08406129
#> 4 0.06714979 0.52719552 0.03335209
#> 5 0.08430487 -0.41755660 0.04590156
#> 6 0.06302876 -0.40071392 0.04181907
plot_pub_bias_compare_3(cmp_mv, main = "Multivariate: publication-bias methods with all-data reference")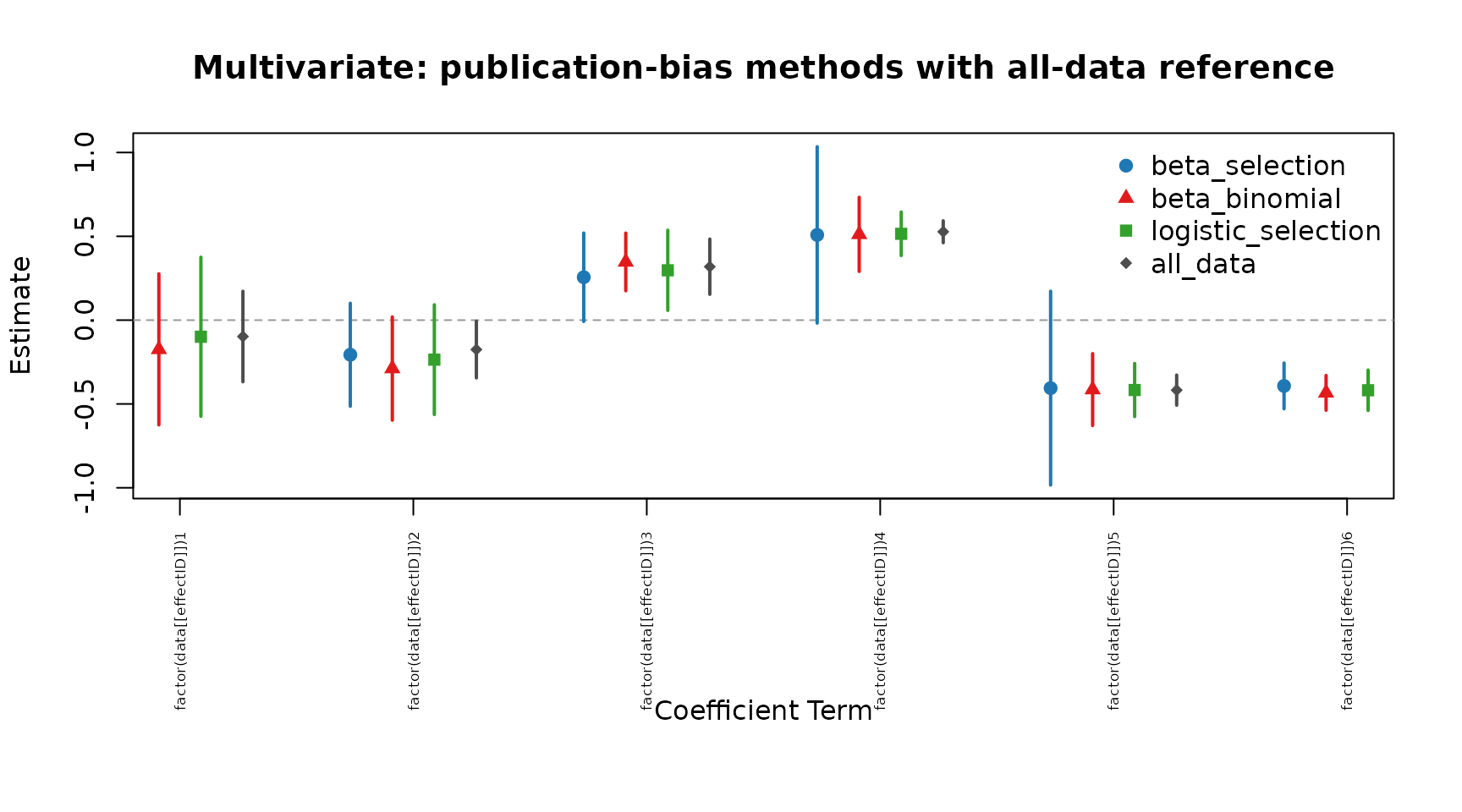
Jackknife version of the multivariate complete-data fit and each publication-bias model:
set.seed(2026)
becker09_jack <- becker09[seq_len(min(6, nrow(becker09))), , drop = FALSE]
jack_mv_complete <- mars_alt_estimation(
type = "jackknife",
data = becker09_jack,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
),
structure = "UN",
estimation_method = "MLE",
jackknife_target = "complete_data"
)
jack_mv_beta_selection <- mars_alt_estimation(
type = "jackknife",
data = becker09_jack,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
),
structure = "UN",
estimation_method = "MLE",
jackknife_target = "beta_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 80,
integration_points = 31
)
)
jack_mv_beta_binomial <- mars_alt_estimation(
type = "jackknife",
data = becker09_jack,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
),
structure = "UN",
estimation_method = "MLE",
jackknife_target = "beta_binomial",
publication_bias_args = list(
p_value_tail = "two.sided",
n_bins = 8,
maxit = 80,
integration_points = 31
)
)
jack_mv_logistic <- mars_alt_estimation(
type = "jackknife",
data = becker09_jack,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
),
structure = "UN",
estimation_method = "MLE",
jackknife_target = "logistic_selection",
publication_bias_args = list(
p_value_tail = "two.sided",
maxit = 80,
integration_points = 31
)
)
jack_mv_complete_stats <- compute_alt_stats(jack_mv_complete, type = "jackknife")
jack_mv_beta_selection_stats <- compute_alt_stats(jack_mv_beta_selection, type = "jackknife")
jack_mv_beta_binomial_stats <- compute_alt_stats(jack_mv_beta_binomial, type = "jackknife")
jack_mv_logistic_stats <- compute_alt_stats(jack_mv_logistic, type = "jackknife")
mv_term <- pb_mv$coefficients$term[1]
jackknife_compare_mv <- rbind(
complete_data = extract_alt_interval(jack_mv_complete_stats, term = mv_term),
beta_selection = extract_alt_interval(jack_mv_beta_selection_stats, term = mv_term),
beta_binomial = extract_alt_interval(jack_mv_beta_binomial_stats, term = mv_term),
logistic_selection = extract_alt_interval(jack_mv_logistic_stats, term = mv_term)
)
jackknife_compare_mvThe multivariate jackknife illustration also uses a small subset so the chunk demonstrates the interface without dominating vignette runtime.
jack_mv_est <- jackknife_compare_mv[, "point_estimate"]
jack_mv_lo <- jackknife_compare_mv[, "ci_lower"]
jack_mv_hi <- jackknife_compare_mv[, "ci_upper"]
xpos_mv <- seq_len(nrow(jackknife_compare_mv))
plot(
xpos_mv, jack_mv_est,
ylim = range(c(jack_mv_lo, jack_mv_hi)),
xaxt = "n", xlab = "", ylab = "Jackknife Estimate",
pch = 19, col = "#1F78B4",
main = paste("Multivariate Jackknife for", mv_term)
)
segments(xpos_mv, jack_mv_lo, xpos_mv, jack_mv_hi, lwd = 2, col = "#1F78B4")
abline(h = 0, lty = 2, col = "gray60")
axis(1, at = xpos_mv, labels = rownames(jackknife_compare_mv), las = 2)
title(xlab = "Jackknife Target", line = 6)Method comparison note: - beta_selection and
beta_binomial use different selection-function
parameterizations. - Coefficients and SEs can differ materially,
especially when beta_binomial uses coarse binning or when
Hessian diagnostics indicate weak identifiability. - The
all_data reference intervals come from the original MARS
fit, while publication-bias intervals come from the publication-bias
likelihood with the conservative Hessian-versus-OPG SE rule. - In
multilevel and multivariate settings, publication-bias fits now retain
the fitted MARS covariance structure through
covariance_mode = "fixed_mars_structure".
The same publication-bias interface can be used across univariate,
multilevel, and multivariate mars objects, and both methods
support funnel plotting.