Synthesis-to-Factor Reporting Workflow
mars authors
2026-05-15
Synthesis-Factor-Report-Workflow.RmdThis vignette demonstrates the new post-synthesis latent functionality and the standardized reporting object:
- Pooled correlation matrix, SE/CI matrix, heterogeneity matrix
- Model decisions (transform, PD repair, missingness handling)
- Factor solution tables (loadings, communalities, fit indices)
- Publication-ready tables
- Plot suite (heatmaps, loadings/path, leave-one-study-out influence)
- Reproducibility and limitations blocks
library(mars)To keep the vignette fast to compile, the live workflow below focuses
on one CFA-based report object. Additional EFA and sensitivity-heavy
variants are shown later as eval = FALSE templates.
1) Fit Correlation Synthesis Model
dat <- na.omit(becker09)
fit_cor <- mars(
data = dat,
studyID = "ID",
effectID = "numID",
sample_size = "N",
effectsize_type = "cor",
varcov_type = "weighted",
variable_names = c(
"Cognitive_Performance",
"Somatic_Performance",
"Selfconfidence_Performance",
"Somatic_Cognitive",
"Selfconfidence_Cognitive",
"Selfconfidence_Somatic"
)
)2) Optional Direct Latent Models
EFA from synthesis
efa_out <- efa_from_synthesis(
mars_object = fit_cor,
n_factors = 1,
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
pd_adjust = "eigen_clip"
)
summary(efa_out)CFA from synthesis
model_cfa <- "
General =~ Performance + Cognitive + Somatic + Selfconfidence
"
rel_vec <- c(
Performance = 0.88,
Cognitive = 0.90,
Somatic = 0.85,
Selfconfidence = 0.87
)
cfa_out <- cfa_from_synthesis(
mars_object = fit_cor,
model = model_cfa,
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
pd_adjust = "eigen_clip"
)
summary(cfa_out)
#> Results generated with MARS:v 0.5.2
#> Friday, May 15, 2026
#>
#> Model Type:
#> multivariate
#>
#> Average Correlation Matrix:
#> Performance Cognitive Somatic Selfconfidence
#> Performance 1.0000000 -0.1194994 -0.1886655 0.3869481
#> Cognitive -0.1194994 1.0000000 0.5232600 -0.4241649
#> Somatic -0.1886655 0.5232600 1.0000000 -0.4225761
#> Selfconfidence 0.3869481 -0.4241649 -0.4225761 1.0000000
#>
#> Synthesis options:
#> method: weighted
#> transform: fisher_z
#> missing_corr: available
#> attenuation: none
#> pd_adjust: eigen_clip
#> pd_adjusted: FALSE
#> min eigen (before/after): 0.46 / 0.46
#> SE note: Latent-model delta SEs are conditional on the synthesized correlation matrix and are not reported. Use se_method = "simulation" or "bootstrap" to propagate correlation uncertainty.
#>
#>
#> Model Fitted:
#>
#> General =~ Performance + Cognitive + Somatic + Selfconfidence
#>
#>
#> Fixed Effects:
#> predictor outcome estimate standard_errors
#> General -> Performance General Performance 1.000000 NA
#> General -> Cognitive General Cognitive -2.098855 NA
#> General -> Somatic General Somatic -2.168363 NA
#> General -> Selfconfidence General Selfconfidence 1.967586 NA
#> test_statistic p_value
#> General -> Performance NA NA
#> General -> Cognitive NA NA
#> General -> Somatic NA NA
#> General -> Selfconfidence NA NA
#>
#>
#> Fit Statistics:
#> Type Value
#> 1 Model Chi-Square 54.499 (2), 0
#> 2 Null Model Chi-Square 432.064 (6)
#> 3 CFI 0.877
#> 4 TLI 0.63
#> 5 RMSEA <NA>
#> 6 SRMR 0.068
#> 7 CFI (raw) 0.877
#> 8 TLI (raw) 0.633) Build Standardized Report Object
CFA report
report_cfa <- synthesis_factor_report(
mars_object = fit_cor,
factor_method = "cfa",
cfa_model = model_cfa,
# group_var = "your_moderator_column", # optional multi-group checks
loading_threshold = 0.30,
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
pd_adjust = "eigen_clip",
include_publication_bias = FALSE
)
report_cfa
#> Synthesis Factor Report
#> Pooled correlation matrix dimension: 4 x 4
#> Factor method: cfa
#> Model decisions:
#> method: weighted
#> transform: fisher_z
#> missing_corr: available
#> attenuation: none
#> pd_adjust: eigen_clip | adjusted: FALSE
#>
#> Fit indices:
#> chi_square df p_value CFI TLI RMSEA SRMR
#> 54.5 2 0 0.8768 0.6303 0.2144 0.06843
#>
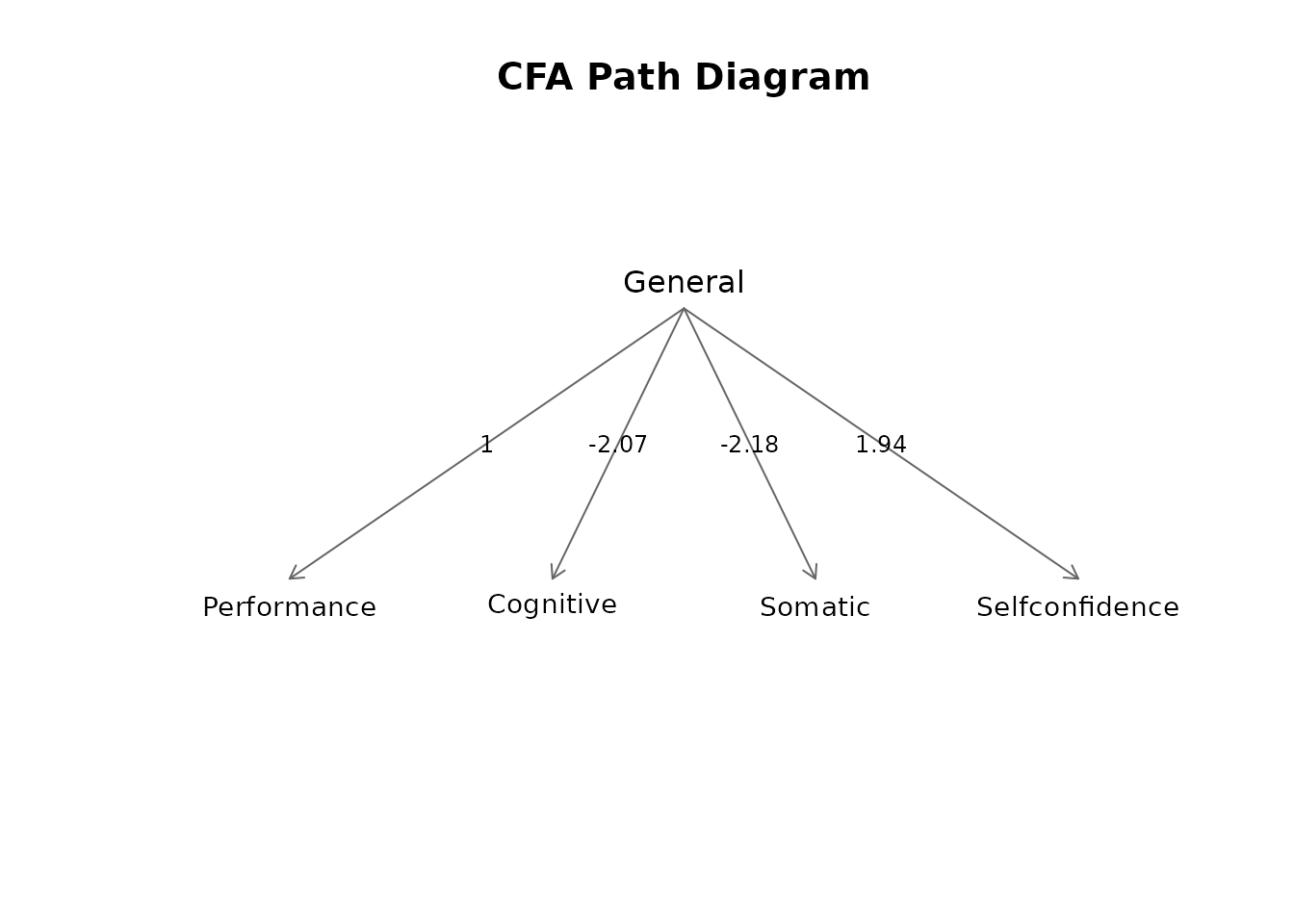
#> Factor loadings table:
#> factor indicator loading salient
#> General Performance 1.000 TRUE
#> General Cognitive -2.099 TRUE
#> General Somatic -2.168 TRUE
#> General Selfconfidence 1.968 TRUE
#>
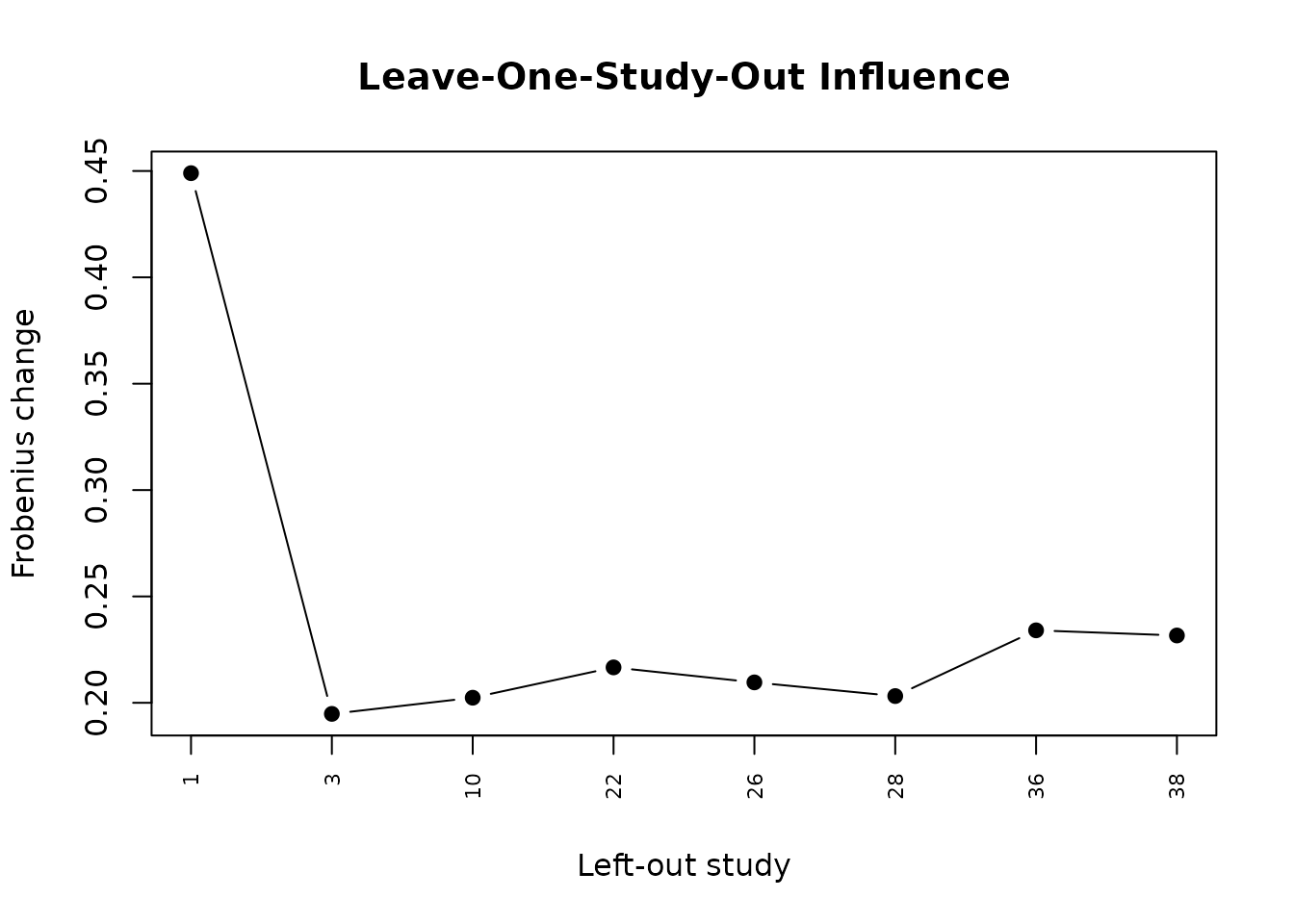
#> Sensitivity:
#> Leave-one-study-out influence was not available.The live report uses a lighter configuration for compilation. More expensive options such as publication-bias checks and sensitivity analyses are shown below as templates.
report_efa <- synthesis_factor_report(
mars_object = fit_cor,
factor_method = "efa",
n_factors = 1,
loading_threshold = 0.30,
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
pd_adjust = "eigen_clip",
include_publication_bias = TRUE
)
report_cfa_sensitivity <- synthesis_factor_report(
mars_object = fit_cor,
factor_method = "cfa",
cfa_model = model_cfa,
loading_threshold = 0.30,
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
missing_corr = "available",
attenuation = "correct",
reliability = rel_vec,
reliability_missing = "impute_mean",
missing_sensitivity = TRUE,
attenuation_sensitivity = TRUE,
reliability_scenarios = list(
lower_rel = c(Performance = 0.80, Cognitive = 0.82, Somatic = 0.78, Selfconfidence = 0.81),
higher_rel = c(Performance = 0.92, Cognitive = 0.94, Somatic = 0.90, Selfconfidence = 0.91)
),
pd_adjust = "eigen_clip",
include_publication_bias = TRUE
)4) Core Outputs
Matrices
report_cfa$pooled_R
#> Performance Cognitive Somatic Selfconfidence
#> Performance 1.0000000 -0.1194994 -0.1886655 0.3869481
#> Cognitive -0.1194994 1.0000000 0.5232600 -0.4241649
#> Somatic -0.1886655 0.5232600 1.0000000 -0.4225761
#> Selfconfidence 0.3869481 -0.4241649 -0.4225761 1.0000000
report_cfa$se_matrix
#> [,1] [,2] [,3] [,4]
#> [1,] 0.00000000 0.13776971 0.08619682 0.08406129
#> [2,] 0.13776971 0.00000000 0.03335209 0.04590156
#> [3,] 0.08619682 0.03335209 0.00000000 0.04181907
#> [4,] 0.08406129 0.04590156 0.04181907 0.00000000
report_cfa$ci_lower_matrix
#> [,1] [,2] [,3] [,4]
#> [1,] 0.0000000 -0.27002863 -0.16894577 -0.16476012
#> [2,] -0.2700286 0.00000000 -0.06537010 -0.08996706
#> [3,] -0.1689458 -0.06537010 0.00000000 -0.08196538
#> [4,] -0.1647601 -0.08996706 -0.08196538 0.00000000
report_cfa$ci_upper_matrix
#> [,1] [,2] [,3] [,4]
#> [1,] 0.0000000 0.27002863 0.16894577 0.16476012
#> [2,] 0.2700286 0.00000000 0.06537010 0.08996706
#> [3,] 0.1689458 0.06537010 0.00000000 0.08196538
#> [4,] 0.1647601 0.08996706 0.08196538 0.00000000
report_cfa$heterogeneity_matrix
#> [,1] [,2] [,3] [,4]
#> [1,] NA 0.36337312 0.21023749 0.20565234
#> [2,] 0.3633731 NA 0.03631344 0.07990099
#> [3,] 0.2102375 0.03631344 NA 0.06352971
#> [4,] 0.2056523 0.07990099 0.06352971 NAModel decisions
report_cfa$model_decisions
#> $synthesis_method
#> [1] "weighted"
#>
#> $synthesis_transform
#> [1] "fisher_z"
#>
#> $missing_corr
#> [1] "available"
#>
#> $attenuation
#> [1] "none"
#>
#> $reliability_missing
#> [1] "error"
#>
#> $pd_adjust
#> [1] "eigen_clip"
#>
#> $pd_adjusted
#> [1] FALSE
#>
#> $missingness_handling
#> NULLFactor solution and fit indices
report_cfa$factor_solution
#> factor indicator loading
#> 1 General Performance 1.000000
#> 2 General Cognitive -2.098855
#> 3 General Somatic -2.168363
#> 4 General Selfconfidence 1.967586
report_cfa$fit_indices
#> chi_square df p_value CFI TLI RMSEA SRMR
#> 1 54.4987 2 0 0.8767819 0.6303458 0.2144082 0.068428045) Publication-Ready Tables
head(report_cfa$publication_tables$pooled_correlation)
#> var1 var2 pooled_r se ci_lower ci_upper
#> 1 Performance Cognitive -0.1194994 0.13776971 -0.27002863 0.27002863
#> 2 Performance Somatic -0.1886655 0.08619682 -0.16894577 0.16894577
#> 3 Cognitive Somatic 0.5232600 0.03335209 -0.06537010 0.06537010
#> 4 Performance Selfconfidence 0.3869481 0.08406129 -0.16476012 0.16476012
#> 5 Cognitive Selfconfidence -0.4241649 0.04590156 -0.08996706 0.08996706
#> 6 Somatic Selfconfidence -0.4225761 0.04181907 -0.08196538 0.08196538
head(report_cfa$publication_tables$heterogeneity)
#> var1 var2 heterogeneity
#> 1 V1 V2 0.36337312
#> 2 V1 V3 0.21023749
#> 3 V2 V3 0.03631344
#> 4 V1 V4 0.20565234
#> 5 V2 V4 0.07990099
#> 6 V3 V4 0.06352971
head(report_cfa$publication_tables$factor_loadings)
#> factor indicator loading salient
#> 1 General Performance 1.000000 TRUE
#> 2 General Cognitive -2.098855 TRUE
#> 3 General Somatic -2.168363 TRUE
#> 4 General Selfconfidence 1.967586 TRUEfactor_loadings includes a salient column
based on the selected loading_threshold.
6) Plot Suite
Heatmaps
plot(report_cfa, type = "pooled_r", main = "Pooled Correlation Heatmap")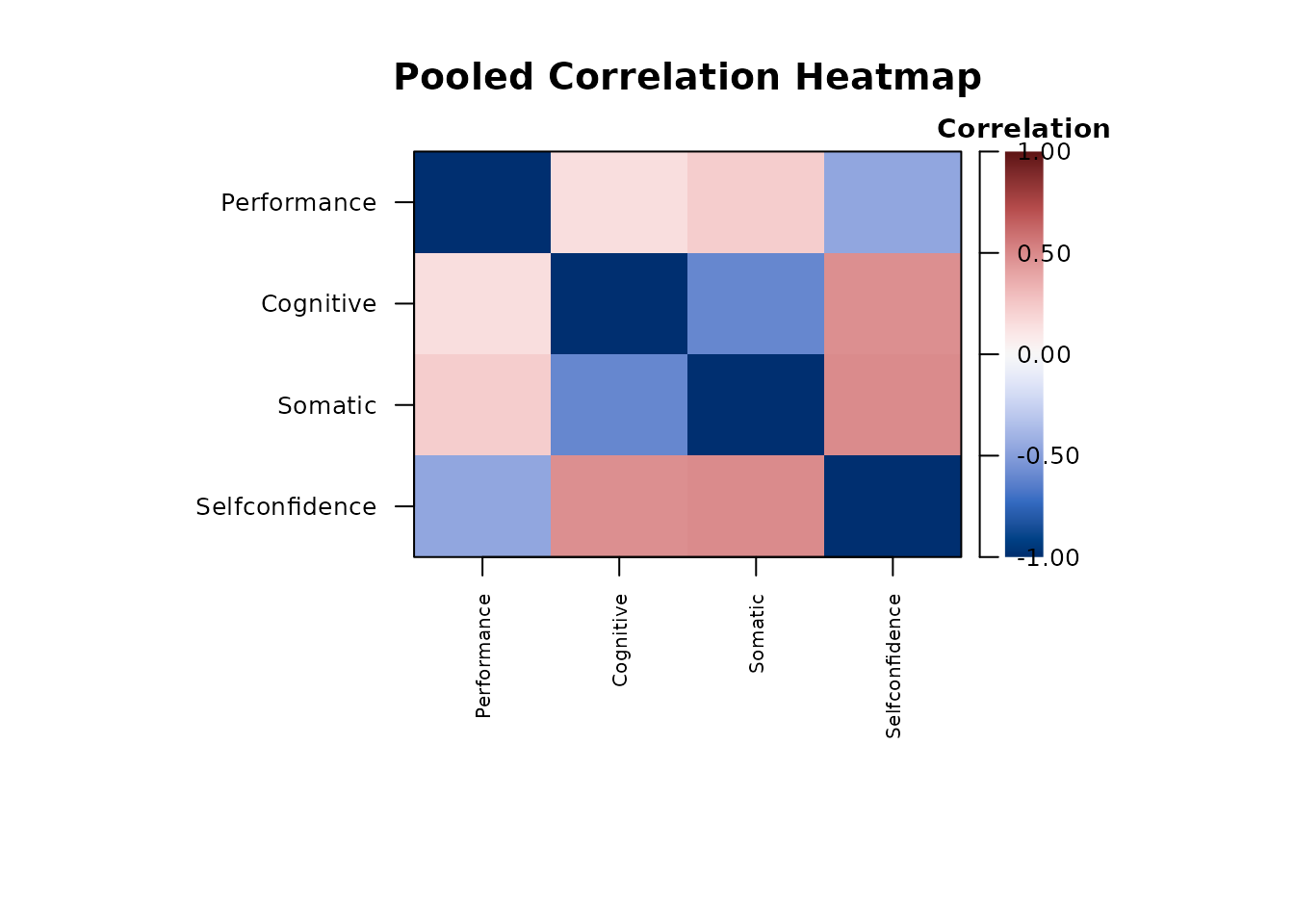
plot(report_cfa, type = "heterogeneity", main = "Heterogeneity Heatmap")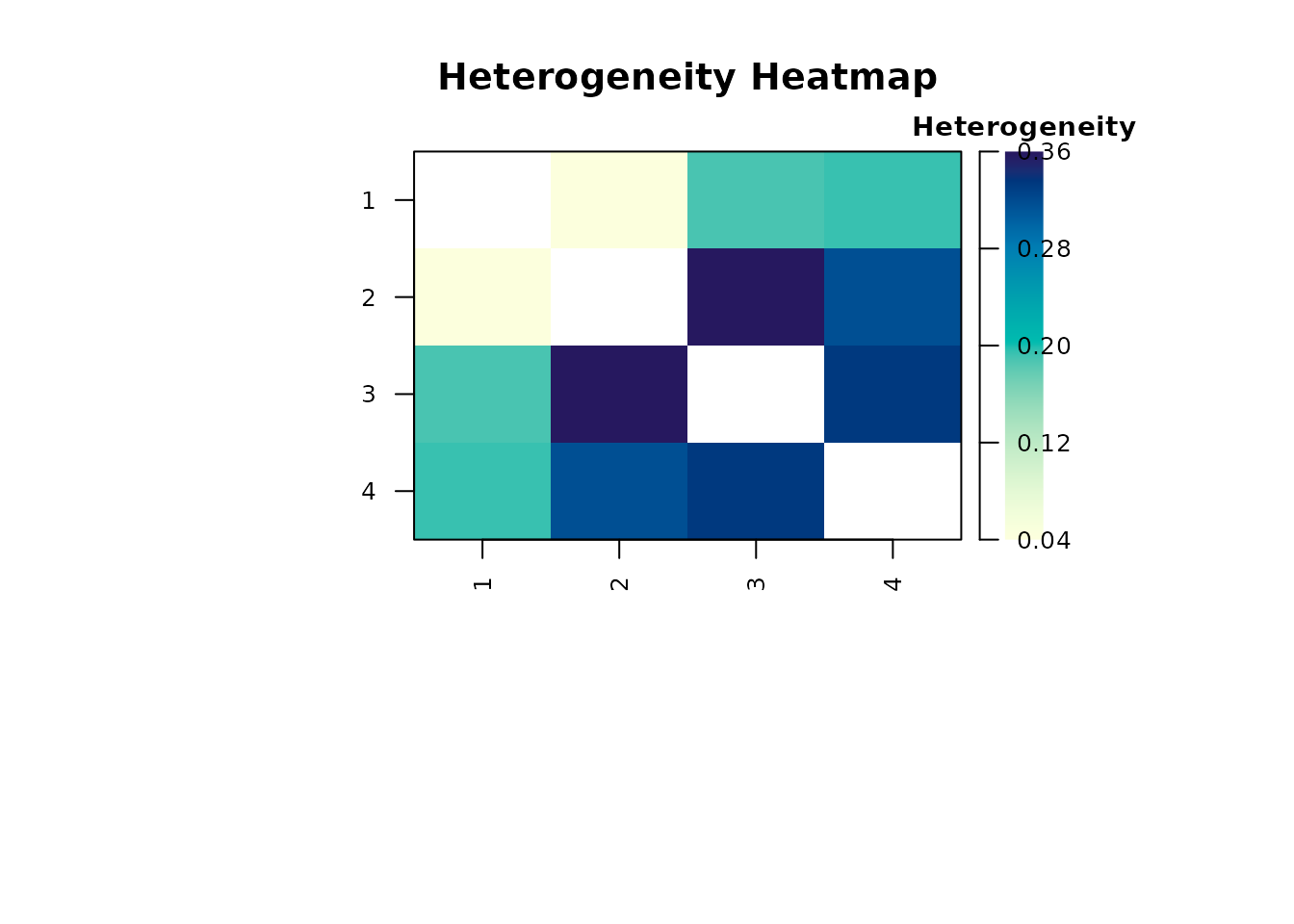
plot(report_cfa, type = "residuals", main = "Residual Heatmap")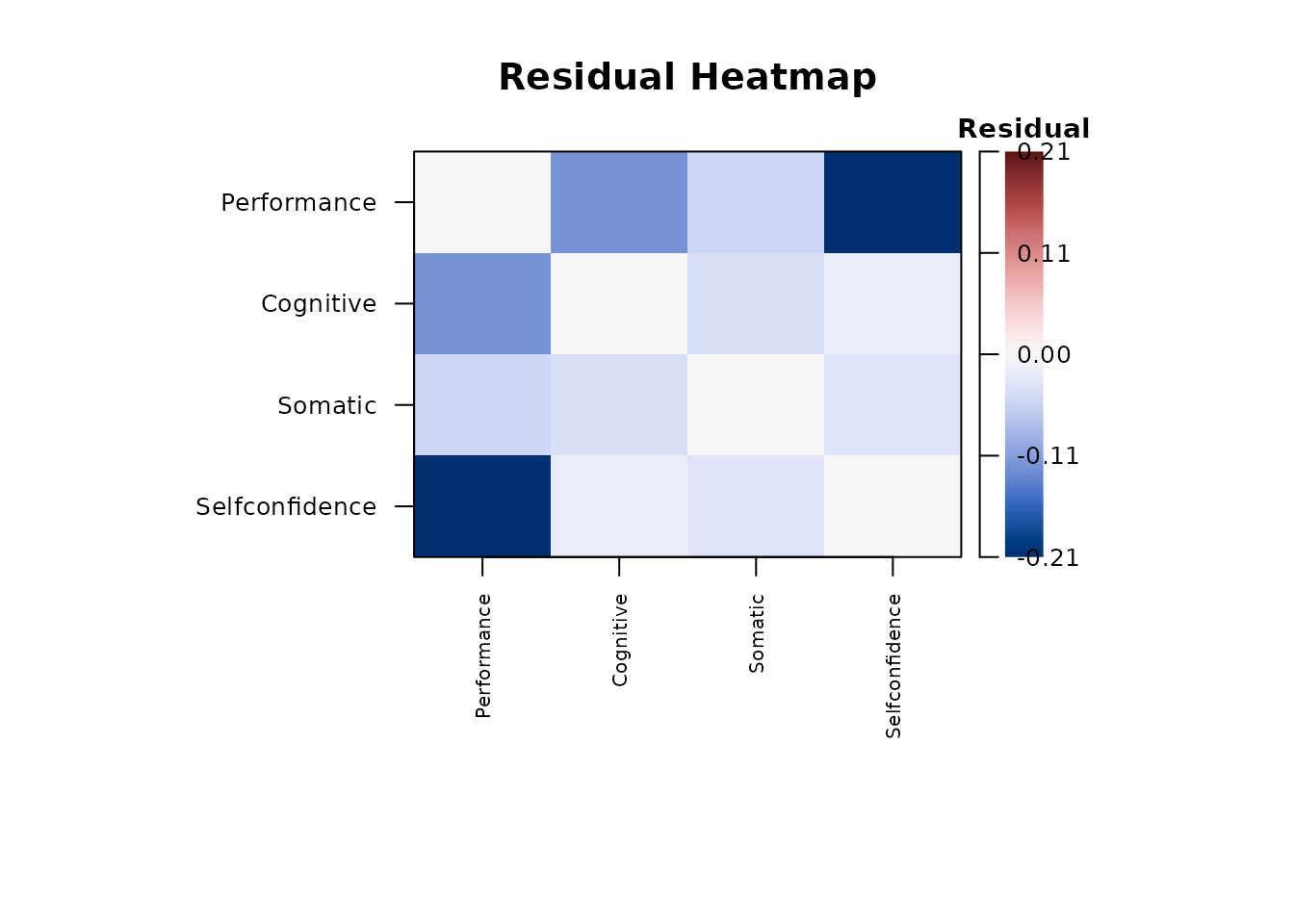
plot(report_cfa, type = "calibration", main = "Calibration Plot")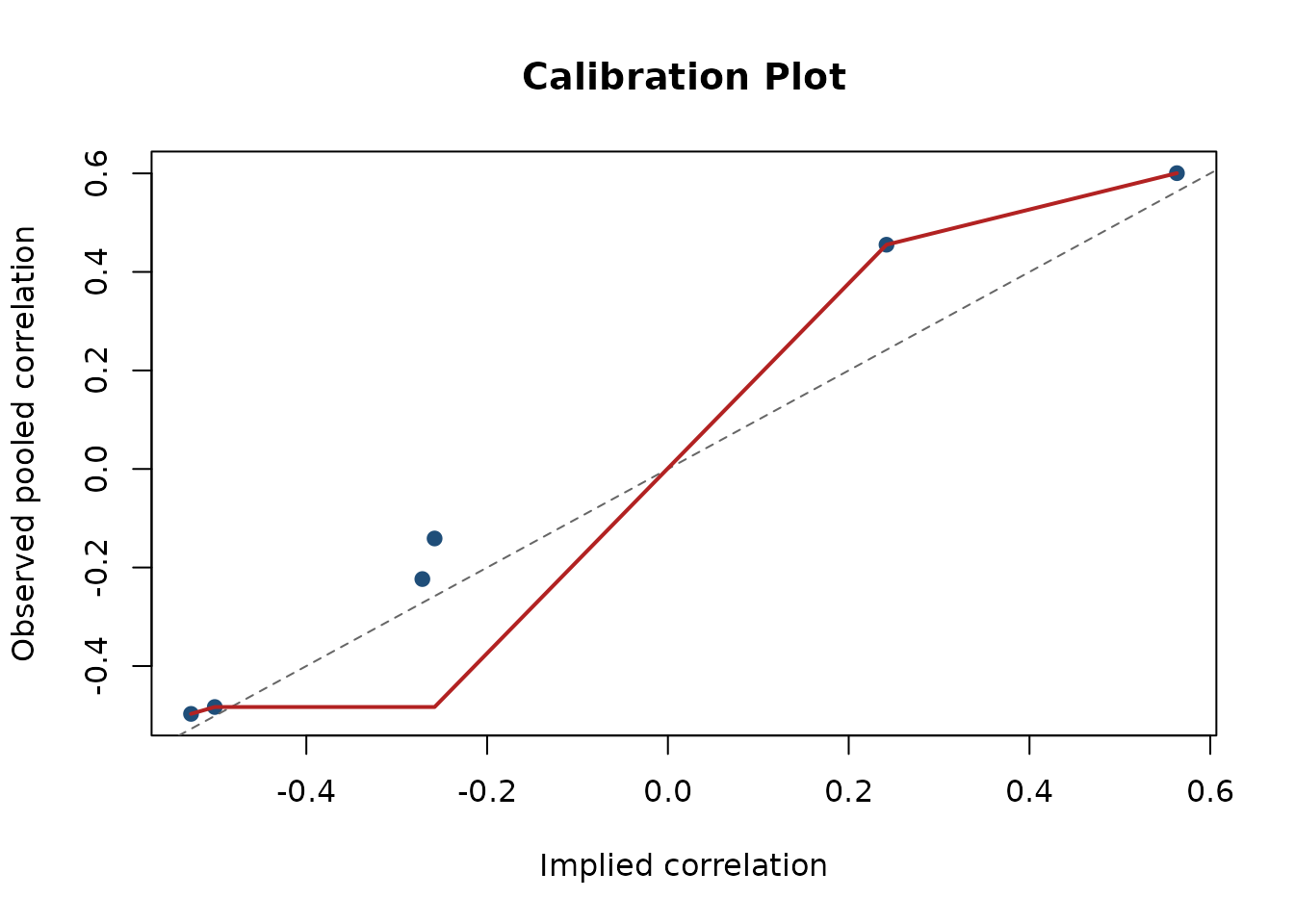
7) Reproducibility Block
report_cfa$reproducibility
#> $call
#> NULL
#>
#> $package_version
#> [1] "0.5.2"
#>
#> $seed
#> [1] 10403
#>
#> $convergence_warnings
#> character(0)This block includes:
- exact call (if stored in the fitted model)
- package version
- seed marker
- convergence warnings (if any)
8) Explicit Limitations Block
report_cfa$limitations
#> $dependence_assumptions
#> [1] "Assumes modeled within-study dependence and random-effects structure are correctly specified for synthesized correlations."
#>
#> $small_study_publication_bias
#> [1] "Not assessed."
#>
#> $sensitivity_results
#> [1] "Leave-one-study-out influence was not available."If missing_sensitivity = TRUE, a missing-correlation
sensitivity block is included:
report_cfa_sensitivity$missing_sensitivity$comparison
report_cfa_sensitivity$attenuation_sensitivity$comparison9) Export Helpers
md_text <- as_markdown(report_cfa)
substr(md_text, 1, 300)
#> [1] "# Synthesis Factor Report\n\n## Model Decisions\n- Synthesis method: `weighted`\n- Transform: `fisher_z`\n- Missing-correlation handling: `available`\n- Attenuation: `none`\n- Reliability missing strategy: `error`\n- PD adjustment: `eigen_clip`\n- Missingness handling: ``\n\n## Fit Indices\n chi_square df p_va"
html_text <- as_html(report_cfa)
substr(html_text, 1, 200)
#> [1] "<html><head><meta charset='utf-8'><title>Synthesis Factor Report</title></head><body><pre># Synthesis Factor Report\n\n## Model Decisions\n- Synthesis method: `weighted`\n- Transform: `fisher_z`\n- Missing"
# DOCX export requires rmarkdown + pandoc:
# as_docx(report_cfa, file = "synthesis-factor-report.docx")10) Optional Multi-Group (Configural vs Metric-Style) Checks
If your synthesized dataset includes a moderator/group variable in
fit_cor$data (for example group_var = "grp"),
you can request configural-vs-metric style checks:
report_group <- synthesis_factor_report(
mars_object = fit_cor,
factor_method = "cfa",
cfa_model = model_cfa,
group_var = "grp",
synthesis_method = "weighted",
synthesis_transform = "fisher_z",
pd_adjust = "eigen_clip"
)
report_group$multi_group$fit_by_group
report_group$multi_group$delta
plot(report_group, type = "invariance")The limitations section explicitly reports:
- dependence assumptions
- small-study/publication-bias check summary
- sensitivity summary from leave-one-study-out diagnostics
11) HSMD Example Data
The package also includes two HSMD example datasets from the
supplementary materials. These can be analyzed directly with
hsmd().
head(hsmd_example_1)
#> Study mT sdT nT mC sdC nC
#> 1 Saw et al. (2019) 68.90 10.29 10 77.80 1.75 10
#> 2 Vuthiarpa et al. (2012) 15.83 4.91 35 20.20 5.86 35
#> 3 Garcia et al. (2013) 12.96 11.30 17 9.49 8.60 19
#> 4 Avci & Kelleci (2016) 12.46 10.38 30 21.90 10.71 30
#> 5 Clark et al. (1995) 17.88 9.30 52 21.67 12.30 68
#> 6 Horowitz et al. (2007) 16.68 10.80 108 20.34 12.60 169
head(hsmd_example_2)
#> Study mT sdT nT mC sdC nC
#> 1 Affleck et al. (1988) 3.13 7.77 39 2.97 8.53 39
#> 2 Bacharach et al. (2010) 17.20 13.28 221 13.90 12.76 99
#> 3 Case-Smith et al. (2014) 28.70 17.15 77 18.50 17.70 55
#> 4 Fontana (2005) 11.20 11.40 17 3.70 8.00 16
#> 5 Jang (2010) 7.15 8.30 28 1.95 7.97 30
#> 6 Marston & Heisted (1994) 9.60 12.90 72 8.60 12.10 62
hsmd_out <- hsmd(
data = hsmd_example_1,
mT = mT, sdT = sdT, nT = nT,
mC = mC, sdC = sdC, nC = nC,
study = Study
)
hsmd_out$summary
#> method est se ci.lb ci.ub tau2 avwW
#> 1 SMD -0.3060830 0.10640132 -0.5146258 -0.09754029 0.11743966 0.04191511
#> 2 HSMD_25 -0.4778418 0.22512747 -0.9190835 -0.03660002 0.72841275 0.04546883
#> 3 HSMD_50 -0.1789429 0.10868797 -0.3919674 0.03408160 0.12560800 0.04186181
#> 4 HSMD_75 -0.1105698 0.08000593 -0.2673785 0.04623900 0.04819645 0.04132897
#> I2
#> 1 60.01581
#> 2 89.56395
#> 3 61.64808
#> 4 38.45138
head(hsmd_out$effect_sizes)
#> d vd g vg d25 vd25 g25
#> 1 -1.2058635 0.23635267 -1.1549116 0.21680118 -0.4859344 0.20590331 -0.4654020
#> 2 -0.8083747 0.06181050 -0.7994259 0.06044957 -0.9824436 0.06403711 -0.9715678
#> 3 0.3483179 0.11314018 0.3405775 0.10816760 1.1779890 0.13072814 1.1518115
#> 4 -0.8951014 0.07334339 -0.8834767 0.07145074 -2.2512745 0.10890197 -2.2220372
#> 5 -0.3413400 0.03442212 -0.3391659 0.03398502 -0.7772932 0.03645409 -0.7723423
#> 6 -0.3067394 0.01534625 -0.3059021 0.01526259 -0.8047337 0.01634537 -0.8025370
#> vg25 d50 vd50 g50 vg50 d75 vd75
#> 1 0.18887064 -0.2340404 0.20136937 -0.2241514 0.18471176 -0.1644206 0.20067585
#> 2 0.06262716 -0.4643390 0.05868293 -0.4591987 0.05739087 -0.3319974 0.05793016
#> 3 0.12498256 0.5997386 0.11645075 0.5864111 0.11133267 0.3999773 0.11367708
#> 4 0.10609172 -1.0166236 0.07527936 -1.0034207 0.07333675 -0.7582966 0.07145845
#> 5 0.03599118 -0.3627068 0.03448480 -0.3603966 0.03404690 -0.2624378 0.03422362
#> 6 0.01625625 -0.3703510 0.01542400 -0.3693400 0.01533991 -0.2714353 0.01530941
#> g75 vg75
#> 1 -0.1574732 0.18407561
#> 2 -0.3283221 0.05665467
#> 3 0.3910889 0.10868090
#> 4 -0.7484486 0.06961444
#> 5 -0.2607662 0.03378904
#> 6 -0.2706943 0.01522594